Chlorine in PDB 7lmw: Receptor For Advanced Glycation End Products VC1 Domain in Complex with 3-(3-((4-(4-Carboxyphenoxy)Benzyl)Oxy)Phenyl)-1H-Indole-2- Carboxylic Acid
Protein crystallography data
The structure of Receptor For Advanced Glycation End Products VC1 Domain in Complex with 3-(3-((4-(4-Carboxyphenoxy)Benzyl)Oxy)Phenyl)-1H-Indole-2- Carboxylic Acid, PDB code: 7lmw
was solved by
L.E.Salay,
N.Kozlyuk,
B.A.Gilston,
R.D.Gogliotti,
P.P.Christov,
K.Kim,
M.Ovee,
A.G.Waterson,
W.J.Chazin,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 33.41 / 2.50 |
| Space group | P 62 |
| Cell size a, b, c (Å), α, β, γ (°) | 101.885, 101.885, 102.306, 90, 90, 120 |
| R / Rfree (%) | 20 / 25.6 |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Receptor For Advanced Glycation End Products VC1 Domain in Complex with 3-(3-((4-(4-Carboxyphenoxy)Benzyl)Oxy)Phenyl)-1H-Indole-2- Carboxylic Acid
(pdb code 7lmw). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total only one binding site of Chlorine was determined in the Receptor For Advanced Glycation End Products VC1 Domain in Complex with 3-(3-((4-(4-Carboxyphenoxy)Benzyl)Oxy)Phenyl)-1H-Indole-2- Carboxylic Acid, PDB code: 7lmw:
In total only one binding site of Chlorine was determined in the Receptor For Advanced Glycation End Products VC1 Domain in Complex with 3-(3-((4-(4-Carboxyphenoxy)Benzyl)Oxy)Phenyl)-1H-Indole-2- Carboxylic Acid, PDB code: 7lmw:
Chlorine binding site 1 out of 1 in 7lmw
Go back to
Chlorine binding site 1 out
of 1 in the Receptor For Advanced Glycation End Products VC1 Domain in Complex with 3-(3-((4-(4-Carboxyphenoxy)Benzyl)Oxy)Phenyl)-1H-Indole-2- Carboxylic Acid
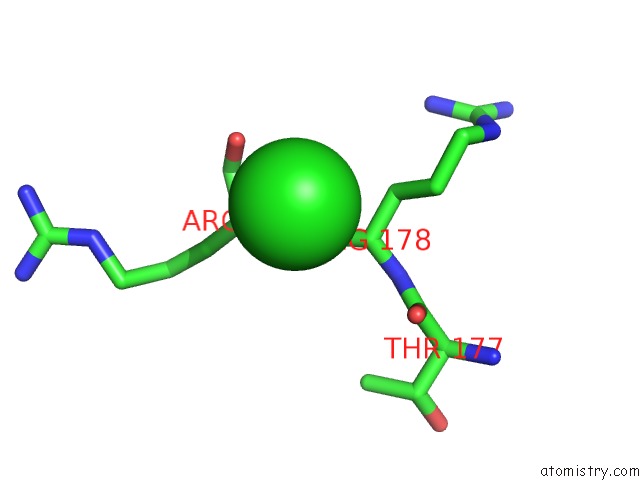
Mono view
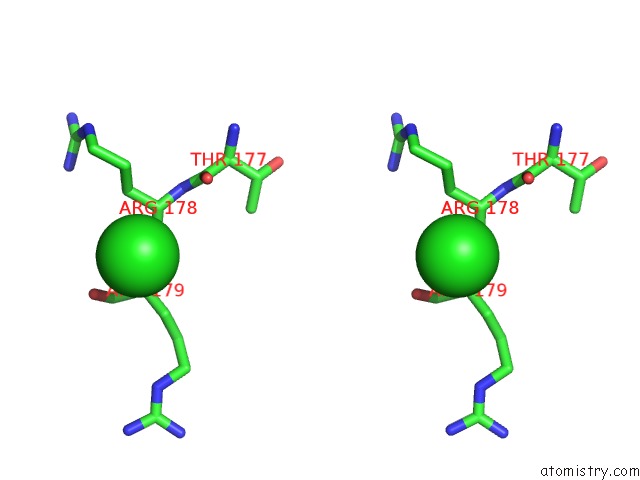
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Receptor For Advanced Glycation End Products VC1 Domain in Complex with 3-(3-((4-(4-Carboxyphenoxy)Benzyl)Oxy)Phenyl)-1H-Indole-2- Carboxylic Acid within 5.0Å range:
|
Reference:
N.Kozlyuk,
B.A.Gilston,
L.E.Salay,
R.D.Gogliotti,
P.P.Christov,
K.Kim,
M.Ovee,
A.G.Waterson,
W.J.Chazin.
A Fragment-Based Approach to Discovery of Receptor For Advanced Glycation End Products Inhibitors. Proteins 2021.
ISSN: ESSN 1097-0134
PubMed: 34156100
DOI: 10.1002/PROT.26162
Page generated: Sat Aug 21 13:21:50 2021
ISSN: ESSN 1097-0134
PubMed: 34156100
DOI: 10.1002/PROT.26162
Last articles
Zn in 8WB0Zn in 8WAX
Zn in 8WAU
Zn in 8WAZ
Zn in 8WAY
Zn in 8WAV
Zn in 8WAW
Zn in 8WAT
Zn in 8W7M
Zn in 8WD3