Chlorine »
PDB 7vc5-7vtv »
7vre »
Chlorine in PDB 7vre: The Crystal Structure of Egfr T790M/C797S with the Inhibitor HCD2892
Enzymatic activity of The Crystal Structure of Egfr T790M/C797S with the Inhibitor HCD2892
All present enzymatic activity of The Crystal Structure of Egfr T790M/C797S with the Inhibitor HCD2892:
2.7.10.1;
2.7.10.1;
Protein crystallography data
The structure of The Crystal Structure of Egfr T790M/C797S with the Inhibitor HCD2892, PDB code: 7vre
was solved by
S.J.Zhu,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 25.85 / 2.51 |
| Space group | I 2 3 |
| Cell size a, b, c (Å), α, β, γ (°) | 146.246, 146.246, 146.246, 90, 90, 90 |
| R / Rfree (%) | 19.9 / 24.2 |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the The Crystal Structure of Egfr T790M/C797S with the Inhibitor HCD2892
(pdb code 7vre). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total 2 binding sites of Chlorine where determined in the The Crystal Structure of Egfr T790M/C797S with the Inhibitor HCD2892, PDB code: 7vre:
Jump to Chlorine binding site number: 1; 2;
In total 2 binding sites of Chlorine where determined in the The Crystal Structure of Egfr T790M/C797S with the Inhibitor HCD2892, PDB code: 7vre:
Jump to Chlorine binding site number: 1; 2;
Chlorine binding site 1 out of 2 in 7vre
Go back to
Chlorine binding site 1 out
of 2 in the The Crystal Structure of Egfr T790M/C797S with the Inhibitor HCD2892
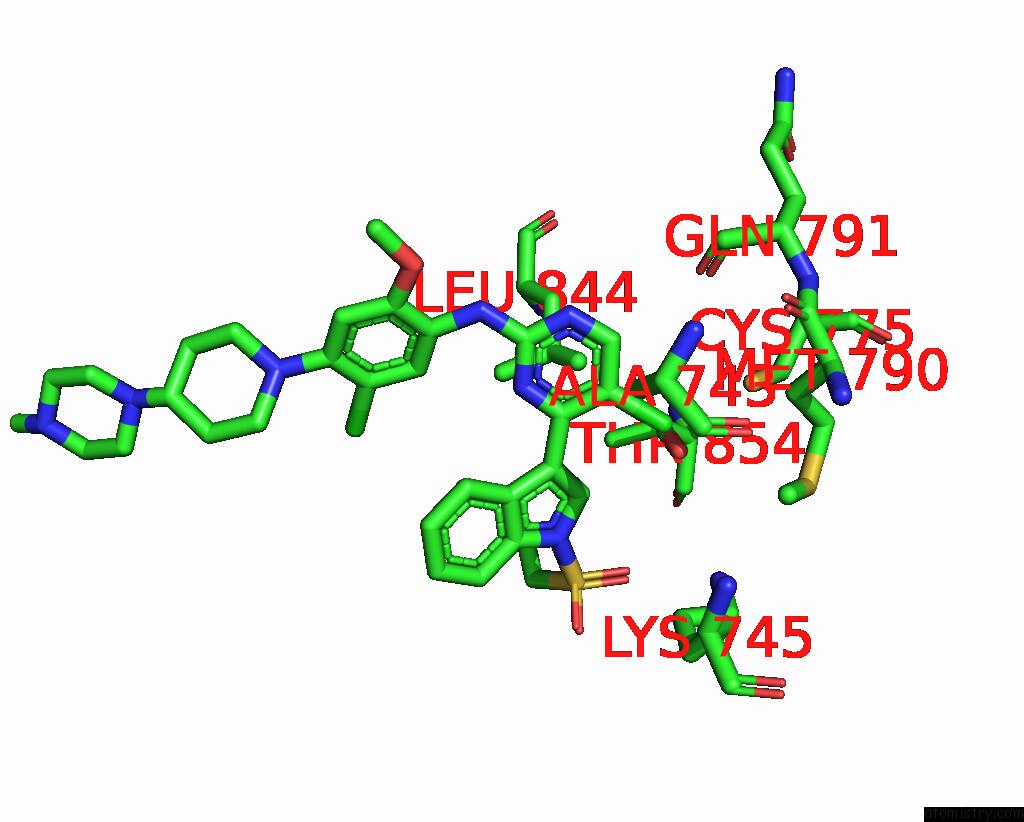
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of The Crystal Structure of Egfr T790M/C797S with the Inhibitor HCD2892 within 5.0Å range:
|
Chlorine binding site 2 out of 2 in 7vre
Go back to
Chlorine binding site 2 out
of 2 in the The Crystal Structure of Egfr T790M/C797S with the Inhibitor HCD2892

Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 2 of The Crystal Structure of Egfr T790M/C797S with the Inhibitor HCD2892 within 5.0Å range:
|
Reference:
H.Chen,
M.Lai,
T.Zhang,
Y.Chen,
L.Tong,
S.Zhu,
Y.Zhou,
X.Ren,
J.Ding,
H.Xie,
X.Lu,
K.Ding.
Conformational Constrained 4-(1-Sulfonyl-3-Indol)Yl-2-Phenylaminopyrimidine Derivatives As New Fourth-Generation Epidermal Growth Factor Receptor Inhibitors Targeting T790M/C797S Mutations. J.Med.Chem. V. 65 6840 2022.
ISSN: ISSN 0022-2623
PubMed: 35446588
DOI: 10.1021/ACS.JMEDCHEM.2C00168
Page generated: Tue Jul 30 05:27:38 2024
ISSN: ISSN 0022-2623
PubMed: 35446588
DOI: 10.1021/ACS.JMEDCHEM.2C00168
Last articles
Zn in 9JPJZn in 9JP7
Zn in 9JPK
Zn in 9JPL
Zn in 9GN6
Zn in 9GN7
Zn in 9GKU
Zn in 9GKW
Zn in 9GKX
Zn in 9GL0