Chlorine in PDB 8gbu: Hepatitis B Capsid Y132A Mutant with Compound Ab-506
Protein crystallography data
The structure of Hepatitis B Capsid Y132A Mutant with Compound Ab-506, PDB code: 8gbu
was solved by
P.S.Horanyi,
S.J.Mayclin,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 97.78 / 2.50 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 151.936, 87.879, 100.827, 90, 104.12, 90 |
| R / Rfree (%) | 23.3 / 28.5 |
Other elements in 8gbu:
The structure of Hepatitis B Capsid Y132A Mutant with Compound Ab-506 also contains other interesting chemical elements:
| Fluorine | (F) | 14 atoms |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Hepatitis B Capsid Y132A Mutant with Compound Ab-506
(pdb code 8gbu). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total 7 binding sites of Chlorine where determined in the Hepatitis B Capsid Y132A Mutant with Compound Ab-506, PDB code: 8gbu:
Jump to Chlorine binding site number: 1; 2; 3; 4; 5; 6; 7;
In total 7 binding sites of Chlorine where determined in the Hepatitis B Capsid Y132A Mutant with Compound Ab-506, PDB code: 8gbu:
Jump to Chlorine binding site number: 1; 2; 3; 4; 5; 6; 7;
Chlorine binding site 1 out of 7 in 8gbu
Go back to
Chlorine binding site 1 out
of 7 in the Hepatitis B Capsid Y132A Mutant with Compound Ab-506

Mono view
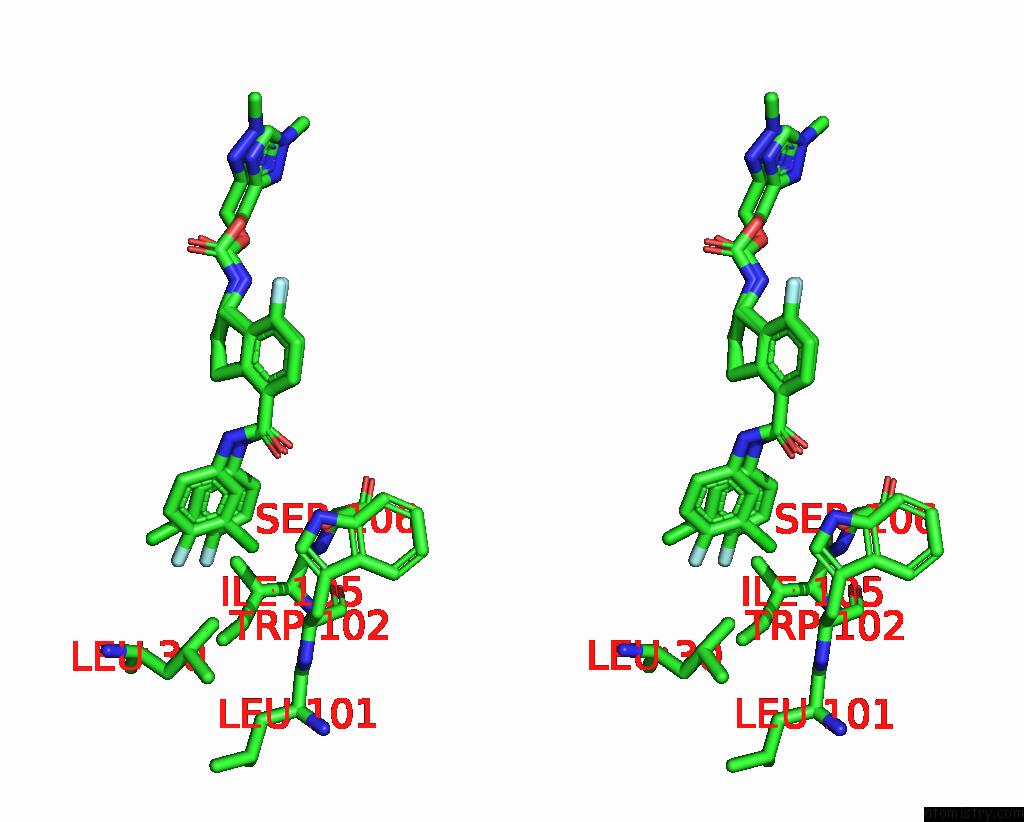
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Hepatitis B Capsid Y132A Mutant with Compound Ab-506 within 5.0Å range:
|
Chlorine binding site 2 out of 7 in 8gbu
Go back to
Chlorine binding site 2 out
of 7 in the Hepatitis B Capsid Y132A Mutant with Compound Ab-506

Mono view
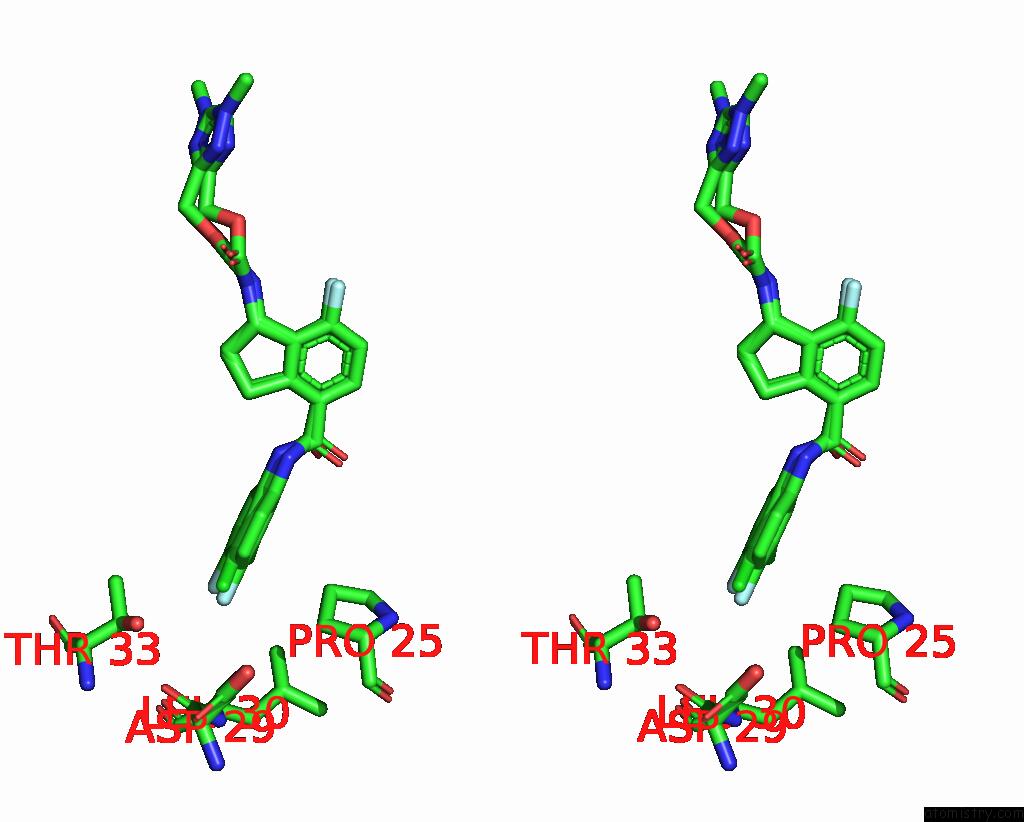
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 2 of Hepatitis B Capsid Y132A Mutant with Compound Ab-506 within 5.0Å range:
|
Chlorine binding site 3 out of 7 in 8gbu
Go back to
Chlorine binding site 3 out
of 7 in the Hepatitis B Capsid Y132A Mutant with Compound Ab-506

Mono view
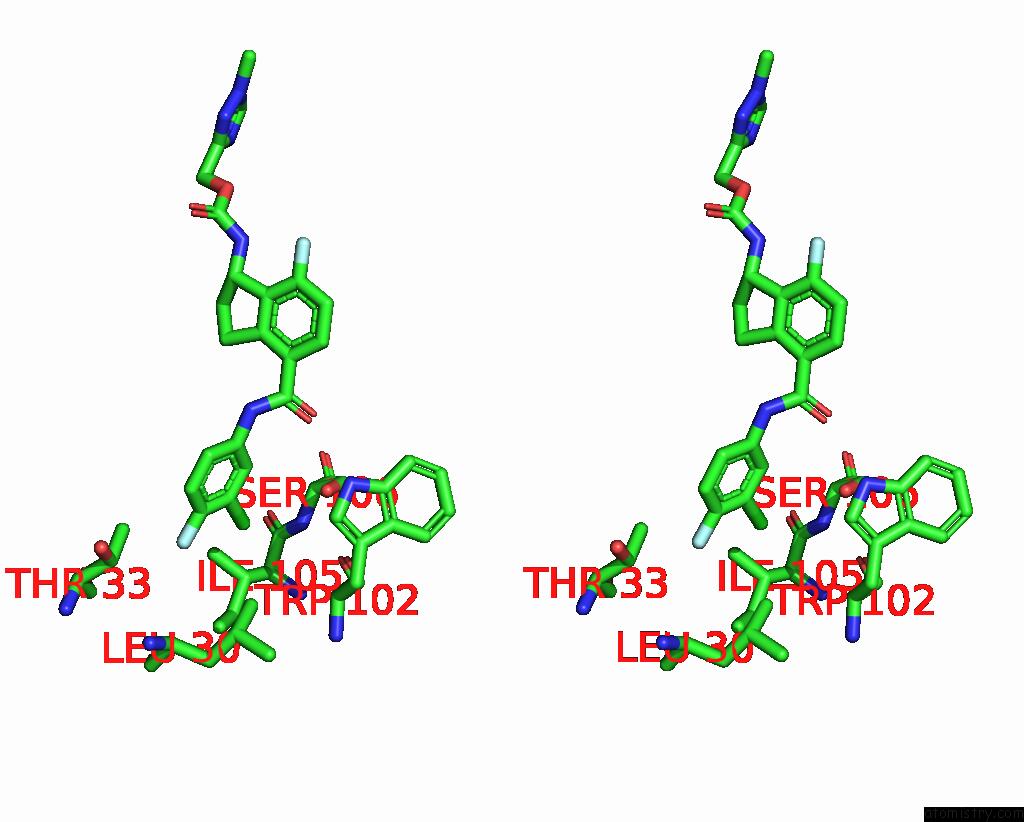
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 3 of Hepatitis B Capsid Y132A Mutant with Compound Ab-506 within 5.0Å range:
|
Chlorine binding site 4 out of 7 in 8gbu
Go back to
Chlorine binding site 4 out
of 7 in the Hepatitis B Capsid Y132A Mutant with Compound Ab-506
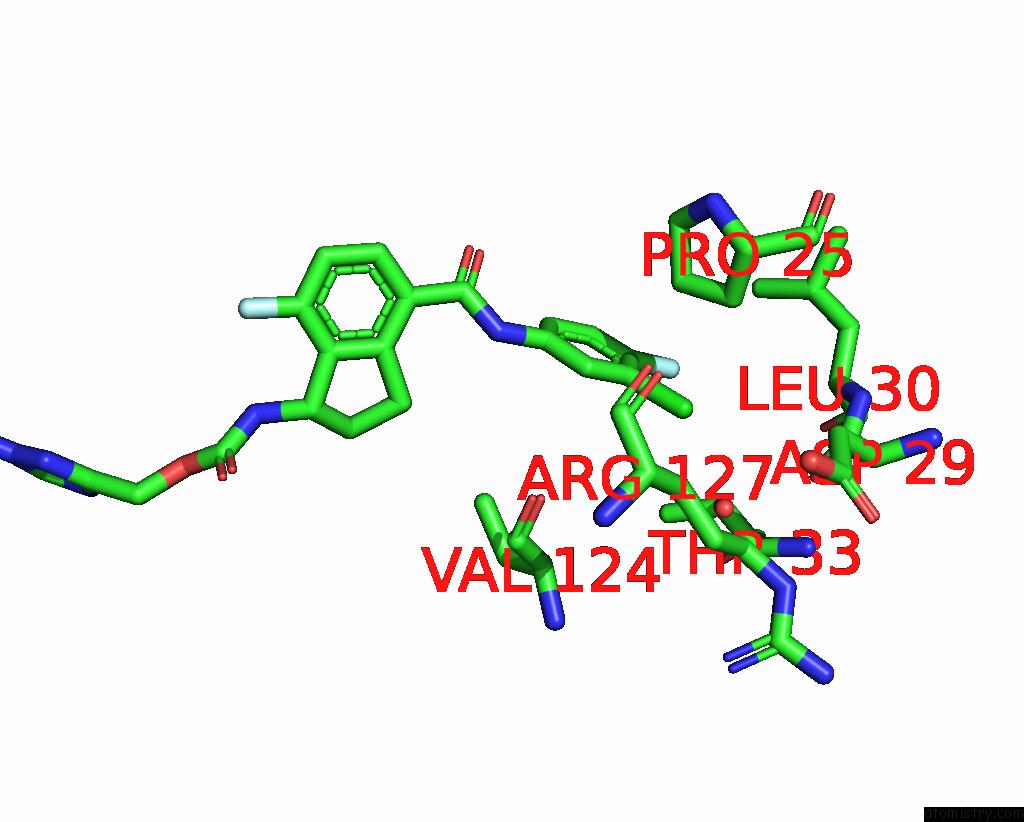
Mono view
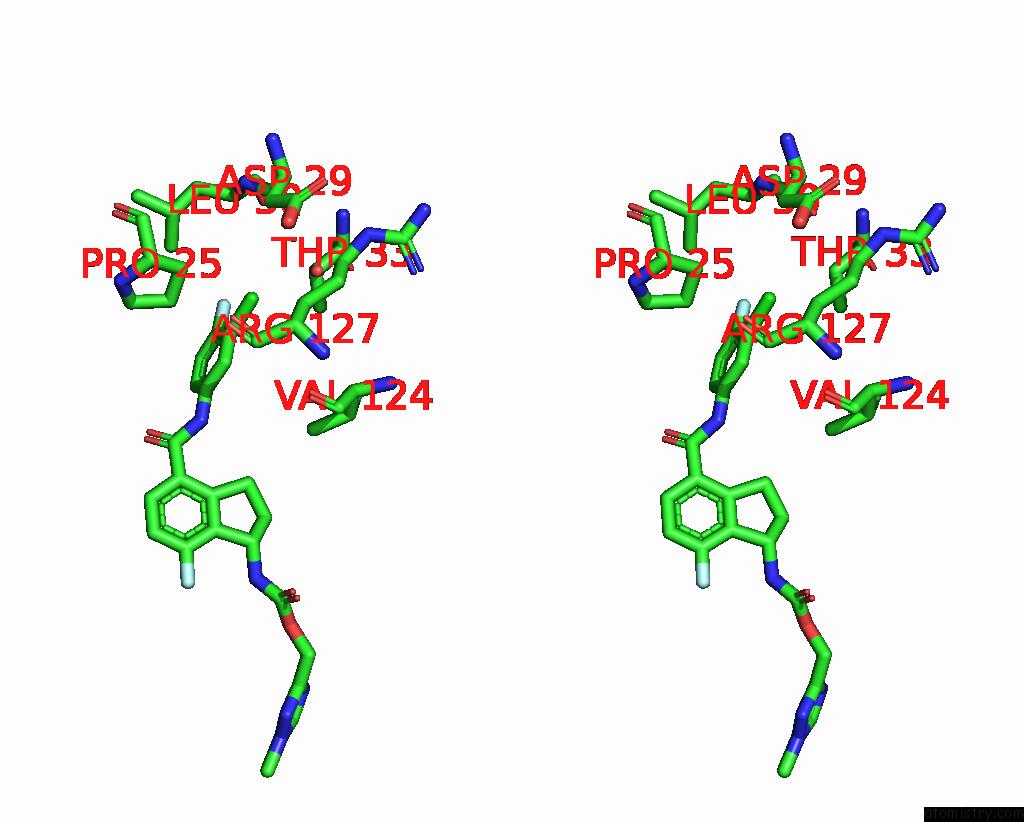
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 4 of Hepatitis B Capsid Y132A Mutant with Compound Ab-506 within 5.0Å range:
|
Chlorine binding site 5 out of 7 in 8gbu
Go back to
Chlorine binding site 5 out
of 7 in the Hepatitis B Capsid Y132A Mutant with Compound Ab-506
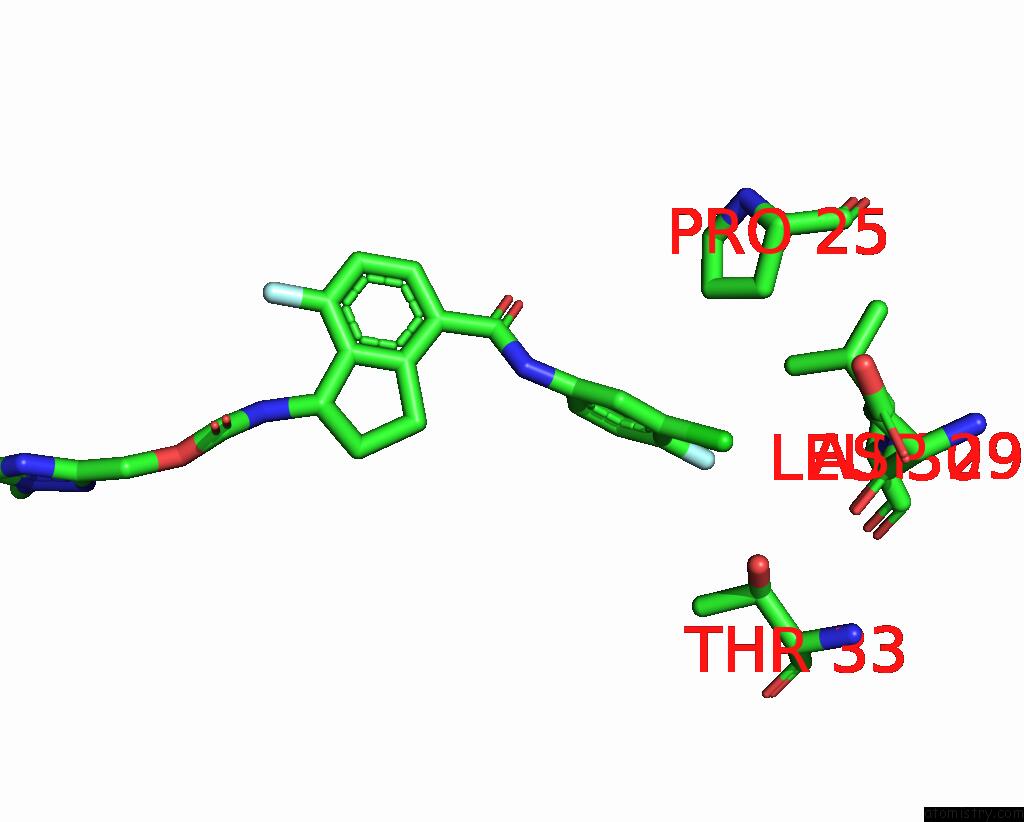
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 5 of Hepatitis B Capsid Y132A Mutant with Compound Ab-506 within 5.0Å range:
|
Chlorine binding site 6 out of 7 in 8gbu
Go back to
Chlorine binding site 6 out
of 7 in the Hepatitis B Capsid Y132A Mutant with Compound Ab-506

Mono view
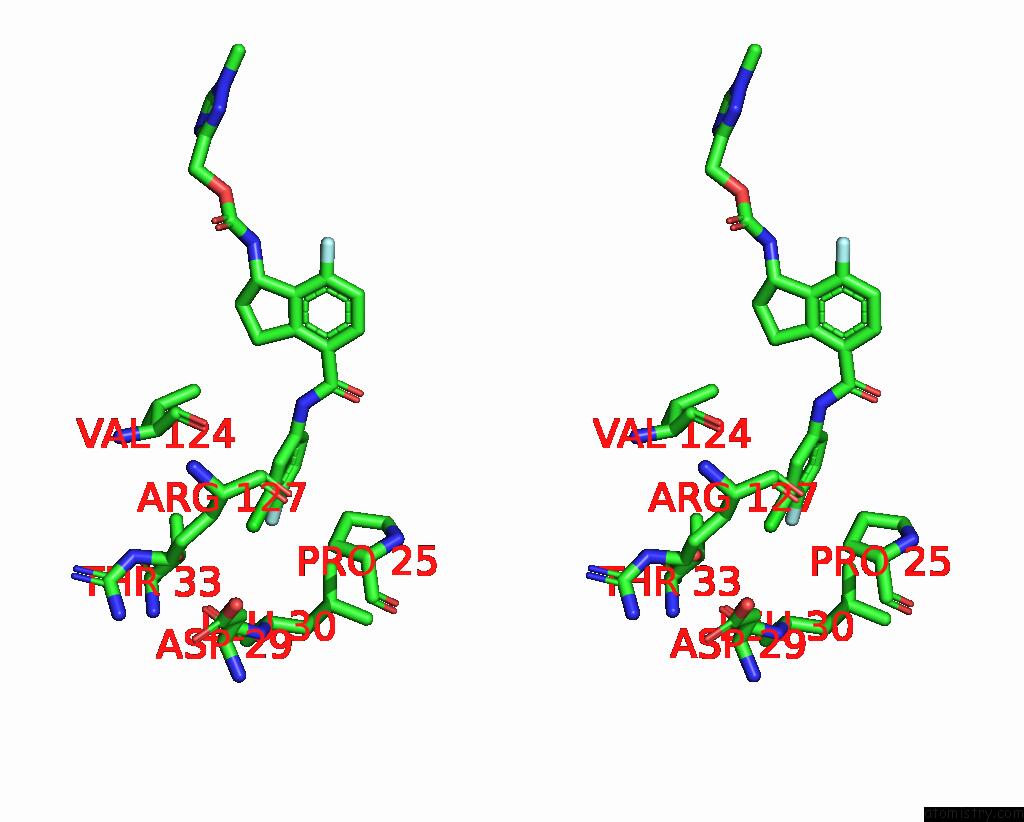
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 6 of Hepatitis B Capsid Y132A Mutant with Compound Ab-506 within 5.0Å range:
|
Chlorine binding site 7 out of 7 in 8gbu
Go back to
Chlorine binding site 7 out
of 7 in the Hepatitis B Capsid Y132A Mutant with Compound Ab-506
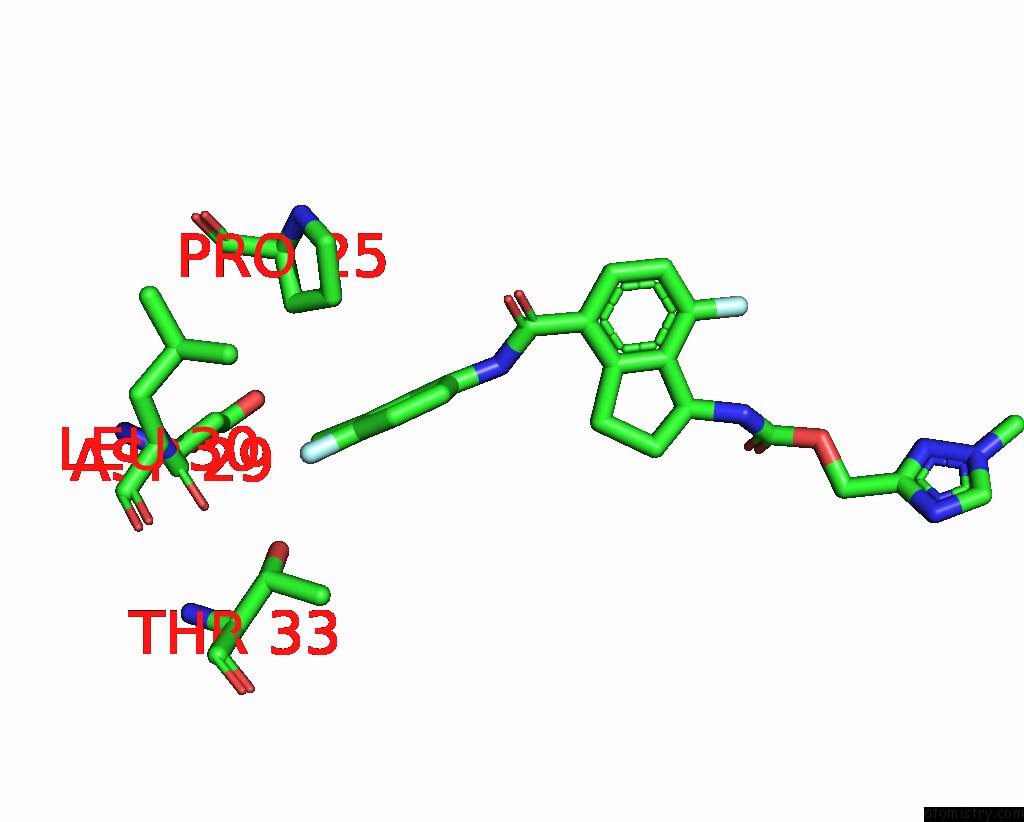
Mono view
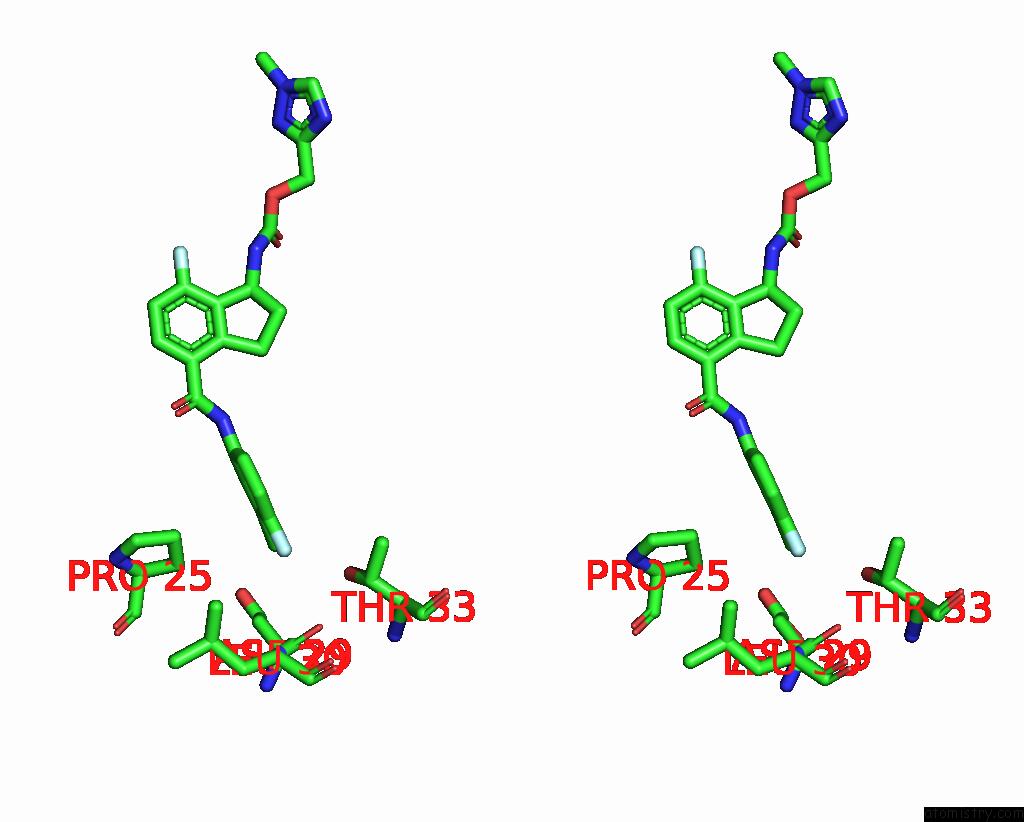
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 7 of Hepatitis B Capsid Y132A Mutant with Compound Ab-506 within 5.0Å range:
|
Reference:
A.G.Cole,
S.G.Kultgen,
N.Mani,
J.G.Quintero,
K.Yi Fan,
A.Ardzinski,
K.Stever,
B.D.Dorsey,
J.R.Phelps,
A.C.H.Lee,
E.P.Thi,
T.Chiu,
S.Tang,
P.S.Horanyi,
S.J.Mayclin,
T.O.Harasym,
M.J.Sofia.
Design, Synthesis, and Structure-Activity Relationship of A Bicyclic Hbv Capsid Assembly Modulator Chemotype Leading to the Identification of Clinical Candidate Ab-506. Bioorg.Med.Chem.Lett. 29456 2023.
ISSN: ESSN 1464-3405
PubMed: 37633618
DOI: 10.1016/J.BMCL.2023.129456
Page generated: Tue Jul 30 09:55:33 2024
ISSN: ESSN 1464-3405
PubMed: 37633618
DOI: 10.1016/J.BMCL.2023.129456
Last articles
Zn in 9MJ5Zn in 9HNW
Zn in 9G0L
Zn in 9FNE
Zn in 9DZN
Zn in 9E0I
Zn in 9D32
Zn in 9DAK
Zn in 8ZXC
Zn in 8ZUF