Chlorine »
PDB 3r41-3rcg »
3r9q »
Chlorine in PDB 3r9q: Structure of A Probable Enoyl-Coa Hydratase/Isomerase From Mycobacterium Abscessus Atcc 19977 / Dsm 44196
Enzymatic activity of Structure of A Probable Enoyl-Coa Hydratase/Isomerase From Mycobacterium Abscessus Atcc 19977 / Dsm 44196
All present enzymatic activity of Structure of A Probable Enoyl-Coa Hydratase/Isomerase From Mycobacterium Abscessus Atcc 19977 / Dsm 44196:
4.2.1.17; 5.3.3.8;
4.2.1.17; 5.3.3.8;
Protein crystallography data
The structure of Structure of A Probable Enoyl-Coa Hydratase/Isomerase From Mycobacterium Abscessus Atcc 19977 / Dsm 44196, PDB code: 3r9q
was solved by
Seattle Structural Genomics Center For Infectious Disease (Ssgcid),
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 50.00 / 2.10 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 98.318, 98.213, 103.951, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 16.1 / 19 |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Structure of A Probable Enoyl-Coa Hydratase/Isomerase From Mycobacterium Abscessus Atcc 19977 / Dsm 44196
(pdb code 3r9q). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total 2 binding sites of Chlorine where determined in the Structure of A Probable Enoyl-Coa Hydratase/Isomerase From Mycobacterium Abscessus Atcc 19977 / Dsm 44196, PDB code: 3r9q:
Jump to Chlorine binding site number: 1; 2;
In total 2 binding sites of Chlorine where determined in the Structure of A Probable Enoyl-Coa Hydratase/Isomerase From Mycobacterium Abscessus Atcc 19977 / Dsm 44196, PDB code: 3r9q:
Jump to Chlorine binding site number: 1; 2;
Chlorine binding site 1 out of 2 in 3r9q
Go back to
Chlorine binding site 1 out
of 2 in the Structure of A Probable Enoyl-Coa Hydratase/Isomerase From Mycobacterium Abscessus Atcc 19977 / Dsm 44196
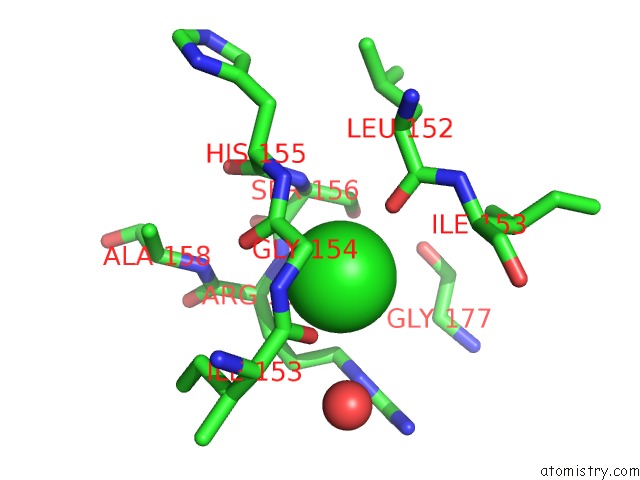
Mono view
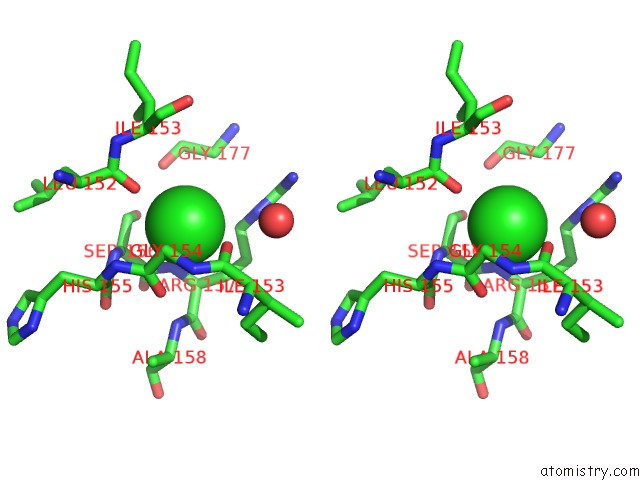
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Structure of A Probable Enoyl-Coa Hydratase/Isomerase From Mycobacterium Abscessus Atcc 19977 / Dsm 44196 within 5.0Å range:
|
Chlorine binding site 2 out of 2 in 3r9q
Go back to
Chlorine binding site 2 out
of 2 in the Structure of A Probable Enoyl-Coa Hydratase/Isomerase From Mycobacterium Abscessus Atcc 19977 / Dsm 44196
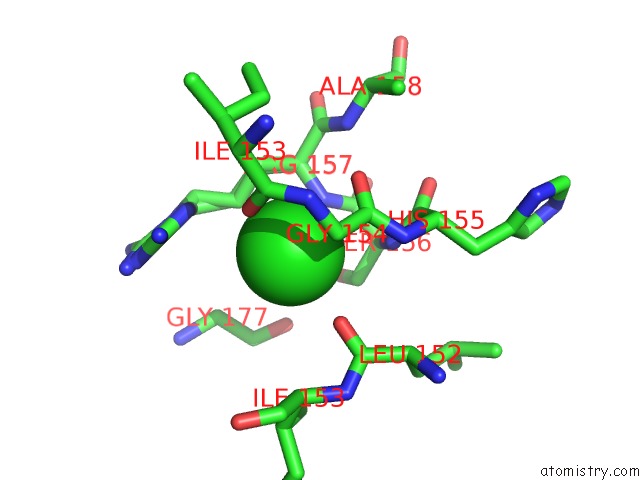
Mono view
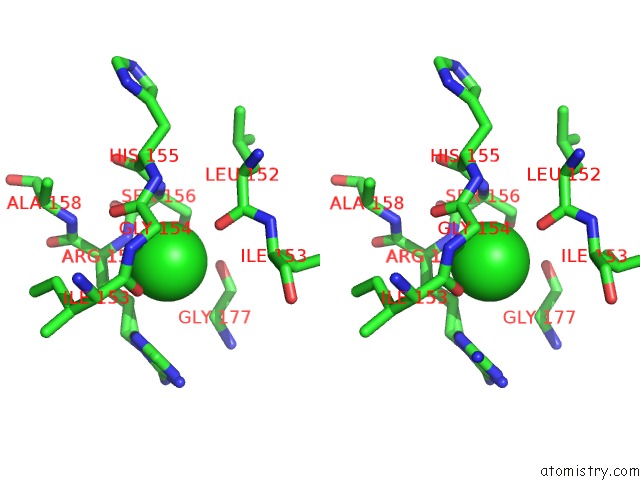
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 2 of Structure of A Probable Enoyl-Coa Hydratase/Isomerase From Mycobacterium Abscessus Atcc 19977 / Dsm 44196 within 5.0Å range:
|
Reference:
L.Baugh,
I.Phan,
D.W.Begley,
M.C.Clifton,
B.Armour,
D.M.Dranow,
B.M.Taylor,
M.M.Muruthi,
J.Abendroth,
J.W.Fairman,
D.Fox,
S.H.Dieterich,
B.L.Staker,
A.S.Gardberg,
R.Choi,
S.N.Hewitt,
A.J.Napuli,
J.Myers,
L.K.Barrett,
Y.Zhang,
M.Ferrell,
E.Mundt,
K.Thompkins,
N.Tran,
S.Lyons-Abbott,
A.Abramov,
A.Sekar,
D.Serbzhinskiy,
D.Lorimer,
G.W.Buchko,
R.Stacy,
L.J.Stewart,
T.E.Edwards,
W.C.Van Voorhis,
P.J.Myler.
Increasing the Structural Coverage of Tuberculosis Drug Targets. Tuberculosis (Edinb) V. 95 142 2015.
ISSN: ISSN 1472-9792
PubMed: 25613812
DOI: 10.1016/J.TUBE.2014.12.003
Page generated: Fri Jul 11 09:45:44 2025
ISSN: ISSN 1472-9792
PubMed: 25613812
DOI: 10.1016/J.TUBE.2014.12.003
Last articles
F in 7RJ8F in 7RH7
F in 7RIV
F in 7RIO
F in 7RIU
F in 7RH0
F in 7RFW
F in 7RH4
F in 7RE3
F in 7RFD