Chlorine »
PDB 3txg-3u69 »
3u0e »
Chlorine in PDB 3u0e: Crystal Structure of Beta-Ketoacyl Synthase From Brucella Melitensis in Complex with Fragment 9320
Enzymatic activity of Crystal Structure of Beta-Ketoacyl Synthase From Brucella Melitensis in Complex with Fragment 9320
All present enzymatic activity of Crystal Structure of Beta-Ketoacyl Synthase From Brucella Melitensis in Complex with Fragment 9320:
2.3.1.41;
2.3.1.41;
Protein crystallography data
The structure of Crystal Structure of Beta-Ketoacyl Synthase From Brucella Melitensis in Complex with Fragment 9320, PDB code: 3u0e
was solved by
Seattle Structural Genomics Center For Infectious Disease (Ssgcid),
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 20.00 / 1.60 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 78.070, 83.750, 73.610, 90.00, 121.50, 90.00 |
| R / Rfree (%) | 13.7 / 15.6 |
Other elements in 3u0e:
The structure of Crystal Structure of Beta-Ketoacyl Synthase From Brucella Melitensis in Complex with Fragment 9320 also contains other interesting chemical elements:
| Sodium | (Na) | 2 atoms |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Crystal Structure of Beta-Ketoacyl Synthase From Brucella Melitensis in Complex with Fragment 9320
(pdb code 3u0e). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total only one binding site of Chlorine was determined in the Crystal Structure of Beta-Ketoacyl Synthase From Brucella Melitensis in Complex with Fragment 9320, PDB code: 3u0e:
In total only one binding site of Chlorine was determined in the Crystal Structure of Beta-Ketoacyl Synthase From Brucella Melitensis in Complex with Fragment 9320, PDB code: 3u0e:
Chlorine binding site 1 out of 1 in 3u0e
Go back to
Chlorine binding site 1 out
of 1 in the Crystal Structure of Beta-Ketoacyl Synthase From Brucella Melitensis in Complex with Fragment 9320
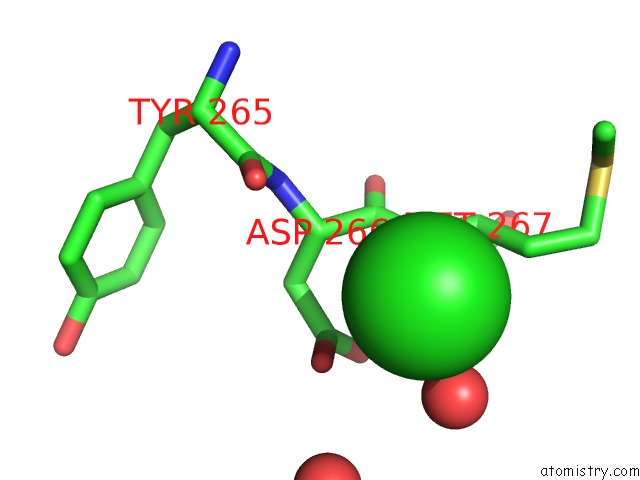
Mono view
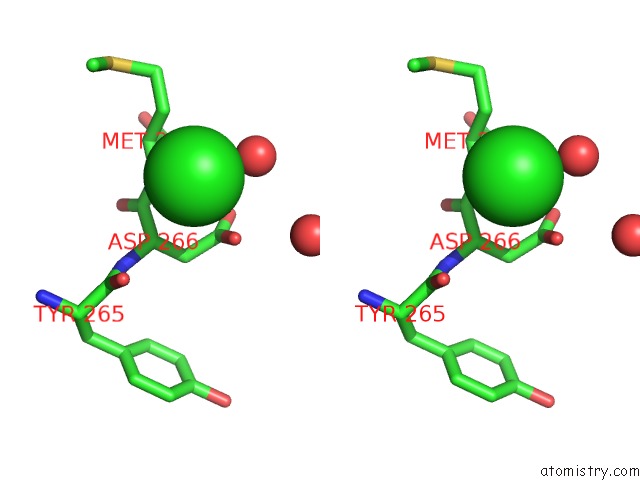
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Crystal Structure of Beta-Ketoacyl Synthase From Brucella Melitensis in Complex with Fragment 9320 within 5.0Å range:
|
Reference:
E.I.Patterson,
J.D.Nanson,
J.Abendroth,
C.Bryan,
B.Sankaran,
P.J.Myler,
J.K.Forwood.
Structural Characterization of Beta-Ketoacyl Acp Synthase I Bound to Platencin and Fragment Screening Molecules at Two Substrate Binding Sites. Proteins 2019.
ISSN: ESSN 1097-0134
PubMed: 31237717
DOI: 10.1002/PROT.25765
Page generated: Sun Jul 21 05:45:19 2024
ISSN: ESSN 1097-0134
PubMed: 31237717
DOI: 10.1002/PROT.25765
Last articles
Zn in 9MJ5Zn in 9HNW
Zn in 9G0L
Zn in 9FNE
Zn in 9DZN
Zn in 9E0I
Zn in 9D32
Zn in 9DAK
Zn in 8ZXC
Zn in 8ZUF