Chlorine »
PDB 4f1b-4fbx »
4fai »
Chlorine in PDB 4fai: Crystal Structure of Mitochondrial Isoform of Glutaminyl Cyclase From Drosophila Melanogaster
Enzymatic activity of Crystal Structure of Mitochondrial Isoform of Glutaminyl Cyclase From Drosophila Melanogaster
All present enzymatic activity of Crystal Structure of Mitochondrial Isoform of Glutaminyl Cyclase From Drosophila Melanogaster:
2.3.2.5;
2.3.2.5;
Protein crystallography data
The structure of Crystal Structure of Mitochondrial Isoform of Glutaminyl Cyclase From Drosophila Melanogaster, PDB code: 4fai
was solved by
P.Kolenko,
B.Koch,
D.Ruiz-Carilo,
M.T.Stubbs,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 71.97 / 1.65 |
| Space group | P 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 46.443, 47.734, 74.556, 85.03, 74.89, 73.90 |
| R / Rfree (%) | 17 / 20.2 |
Other elements in 4fai:
The structure of Crystal Structure of Mitochondrial Isoform of Glutaminyl Cyclase From Drosophila Melanogaster also contains other interesting chemical elements:
| Zinc | (Zn) | 2 atoms |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Crystal Structure of Mitochondrial Isoform of Glutaminyl Cyclase From Drosophila Melanogaster
(pdb code 4fai). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total 2 binding sites of Chlorine where determined in the Crystal Structure of Mitochondrial Isoform of Glutaminyl Cyclase From Drosophila Melanogaster, PDB code: 4fai:
Jump to Chlorine binding site number: 1; 2;
In total 2 binding sites of Chlorine where determined in the Crystal Structure of Mitochondrial Isoform of Glutaminyl Cyclase From Drosophila Melanogaster, PDB code: 4fai:
Jump to Chlorine binding site number: 1; 2;
Chlorine binding site 1 out of 2 in 4fai
Go back to
Chlorine binding site 1 out
of 2 in the Crystal Structure of Mitochondrial Isoform of Glutaminyl Cyclase From Drosophila Melanogaster
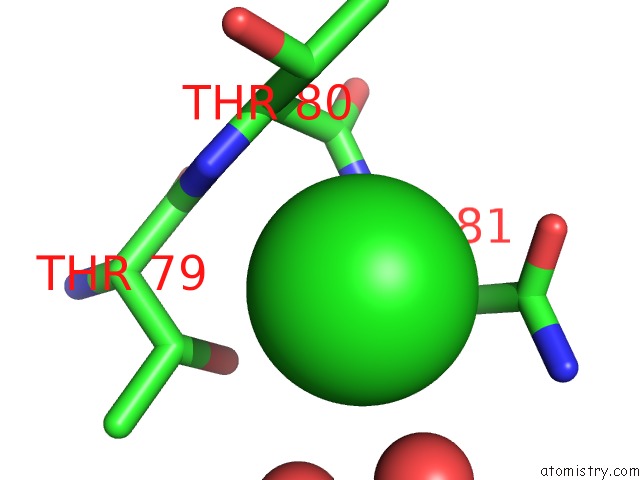
Mono view
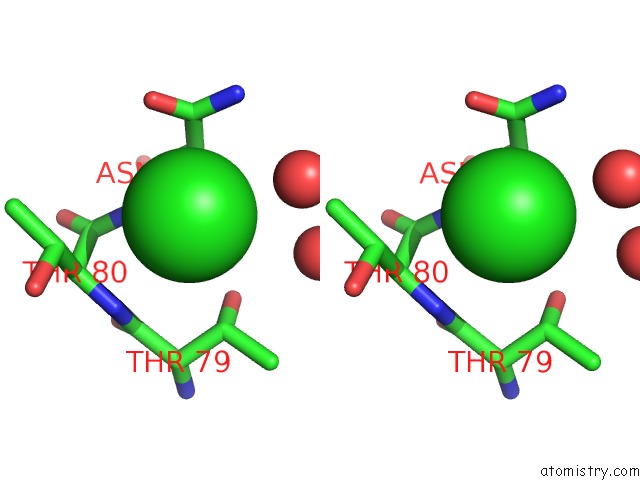
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Crystal Structure of Mitochondrial Isoform of Glutaminyl Cyclase From Drosophila Melanogaster within 5.0Å range:
|
Chlorine binding site 2 out of 2 in 4fai
Go back to
Chlorine binding site 2 out
of 2 in the Crystal Structure of Mitochondrial Isoform of Glutaminyl Cyclase From Drosophila Melanogaster
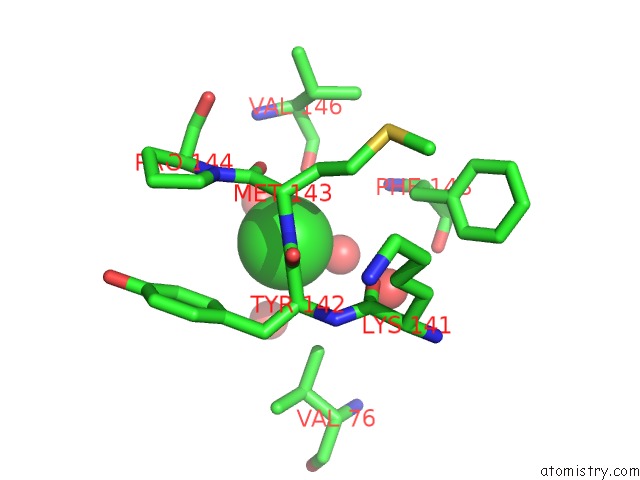
Mono view
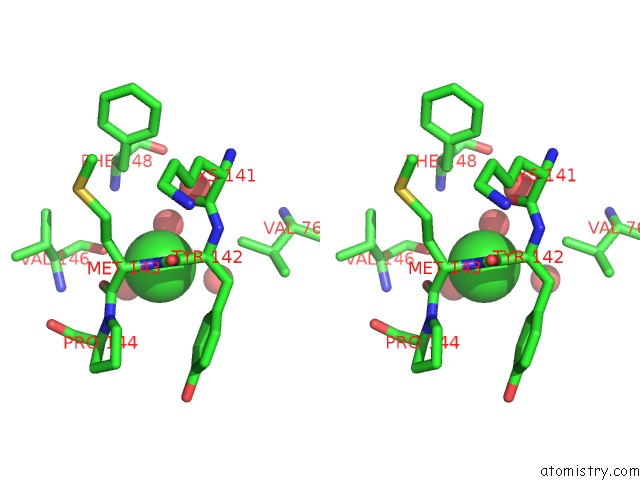
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 2 of Crystal Structure of Mitochondrial Isoform of Glutaminyl Cyclase From Drosophila Melanogaster within 5.0Å range:
|
Reference:
B.Koch,
P.Kolenko,
M.Buchholz,
D.Ruiz Carrillo,
C.Parthier,
M.Wermann,
J.U.Rahfeld,
G.Reuter,
S.Schilling,
M.T.Stubbs,
H.U.Demuth.
Crystal Structures of Glutaminyl Cyclases (Qcs) From Drosophila Melanogaster Reveal Active Site Conservation Between Insect and Mammalian Qcs. Biochemistry V. 51 7383 2012.
ISSN: ISSN 0006-2960
PubMed: 22897232
DOI: 10.1021/BI300687G
Page generated: Fri Jul 11 15:09:37 2025
ISSN: ISSN 0006-2960
PubMed: 22897232
DOI: 10.1021/BI300687G
Last articles
Cl in 7ZSGCl in 7ZNM
Cl in 7ZSF
Cl in 7ZQX
Cl in 7ZR4
Cl in 7ZR3
Cl in 7ZQQ
Cl in 7ZQI
Cl in 7ZPG
Cl in 7ZL7