Chlorine »
PDB 4lxh-4m5g »
4m54 »
Chlorine in PDB 4m54: The Structure of the Staphyloferrin B Precursor Biosynthetic Enzyme Sbnb Bound to N-(1-Amino-1-Carboxyl-2-Ethyl)-Glutamic Acid and Nadh
Protein crystallography data
The structure of The Structure of the Staphyloferrin B Precursor Biosynthetic Enzyme Sbnb Bound to N-(1-Amino-1-Carboxyl-2-Ethyl)-Glutamic Acid and Nadh, PDB code: 4m54
was solved by
M.J.Kobylarz,
M.E.P.Murphy,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 29.73 / 2.36 |
| Space group | P 41 21 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 63.116, 63.116, 159.413, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 22.3 / 25.8 |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the The Structure of the Staphyloferrin B Precursor Biosynthetic Enzyme Sbnb Bound to N-(1-Amino-1-Carboxyl-2-Ethyl)-Glutamic Acid and Nadh
(pdb code 4m54). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total only one binding site of Chlorine was determined in the The Structure of the Staphyloferrin B Precursor Biosynthetic Enzyme Sbnb Bound to N-(1-Amino-1-Carboxyl-2-Ethyl)-Glutamic Acid and Nadh, PDB code: 4m54:
In total only one binding site of Chlorine was determined in the The Structure of the Staphyloferrin B Precursor Biosynthetic Enzyme Sbnb Bound to N-(1-Amino-1-Carboxyl-2-Ethyl)-Glutamic Acid and Nadh, PDB code: 4m54:
Chlorine binding site 1 out of 1 in 4m54
Go back to
Chlorine binding site 1 out
of 1 in the The Structure of the Staphyloferrin B Precursor Biosynthetic Enzyme Sbnb Bound to N-(1-Amino-1-Carboxyl-2-Ethyl)-Glutamic Acid and Nadh
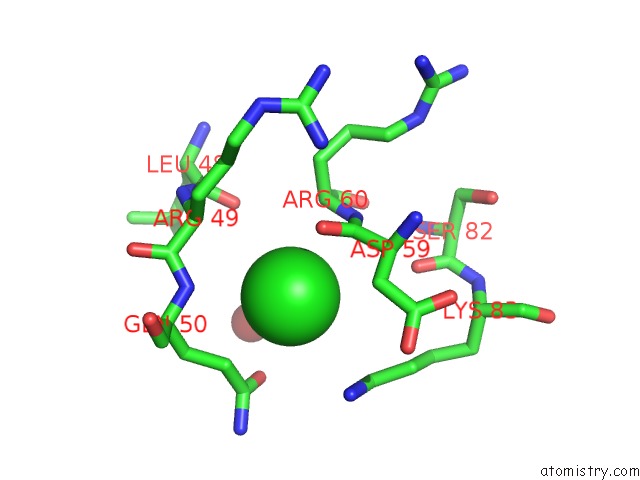
Mono view
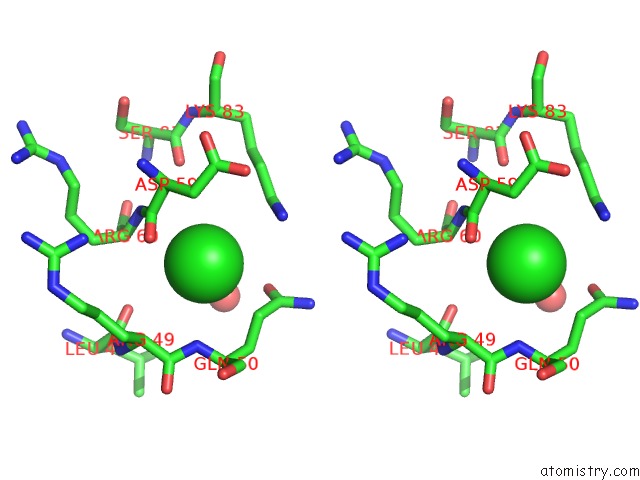
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of The Structure of the Staphyloferrin B Precursor Biosynthetic Enzyme Sbnb Bound to N-(1-Amino-1-Carboxyl-2-Ethyl)-Glutamic Acid and Nadh within 5.0Å range:
|
Reference:
M.J.Kobylarz,
J.C.Grigg,
S.J.Takayama,
D.K.Rai,
D.E.Heinrichs,
M.E.Murphy.
Synthesis of L-2,3-Diaminopropionic Acid, A Siderophore and Antibiotic Precursor. Chem.Biol. V. 21 379 2014.
ISSN: ISSN 1074-5521
PubMed: 24485762
DOI: 10.1016/J.CHEMBIOL.2013.12.011
Page generated: Sun Jul 21 19:34:08 2024
ISSN: ISSN 1074-5521
PubMed: 24485762
DOI: 10.1016/J.CHEMBIOL.2013.12.011
Last articles
Ca in 5NERCa in 5NEM
Ca in 5NE5
Ca in 5NBP
Ca in 5NBN
Ca in 5NBM
Ca in 5NBL
Ca in 5N7G
Ca in 5N7F
Ca in 5N7D