Chlorine »
PDB 4qz1-4r69 »
4qzs »
Chlorine in PDB 4qzs: Crystal Structure of the First Bromodomain of Human 3-Fluoro Tyrosine- Labeled BRD4 in Complex with JQ1
Protein crystallography data
The structure of Crystal Structure of the First Bromodomain of Human 3-Fluoro Tyrosine- Labeled BRD4 in Complex with JQ1, PDB code: 4qzs
was solved by
S.W.Ember,
E.Schonbrunn,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 19.65 / 1.45 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 36.767, 44.898, 78.597, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 14 / 18.5 |
Other elements in 4qzs:
The structure of Crystal Structure of the First Bromodomain of Human 3-Fluoro Tyrosine- Labeled BRD4 in Complex with JQ1 also contains other interesting chemical elements:
| Fluorine | (F) | 11 atoms |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Crystal Structure of the First Bromodomain of Human 3-Fluoro Tyrosine- Labeled BRD4 in Complex with JQ1
(pdb code 4qzs). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total 2 binding sites of Chlorine where determined in the Crystal Structure of the First Bromodomain of Human 3-Fluoro Tyrosine- Labeled BRD4 in Complex with JQ1, PDB code: 4qzs:
Jump to Chlorine binding site number: 1; 2;
In total 2 binding sites of Chlorine where determined in the Crystal Structure of the First Bromodomain of Human 3-Fluoro Tyrosine- Labeled BRD4 in Complex with JQ1, PDB code: 4qzs:
Jump to Chlorine binding site number: 1; 2;
Chlorine binding site 1 out of 2 in 4qzs
Go back to
Chlorine binding site 1 out
of 2 in the Crystal Structure of the First Bromodomain of Human 3-Fluoro Tyrosine- Labeled BRD4 in Complex with JQ1
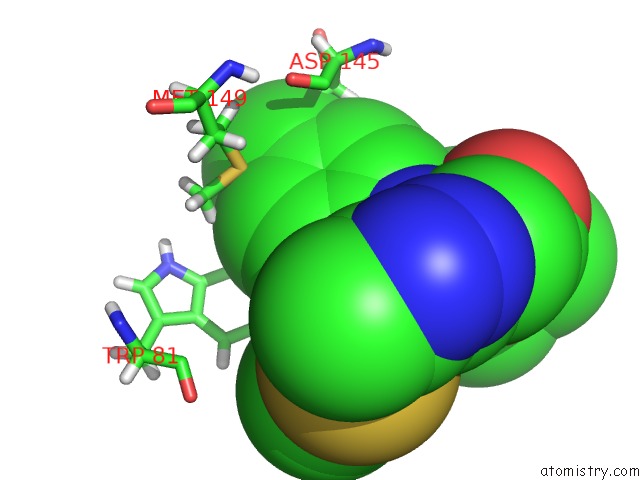
Mono view
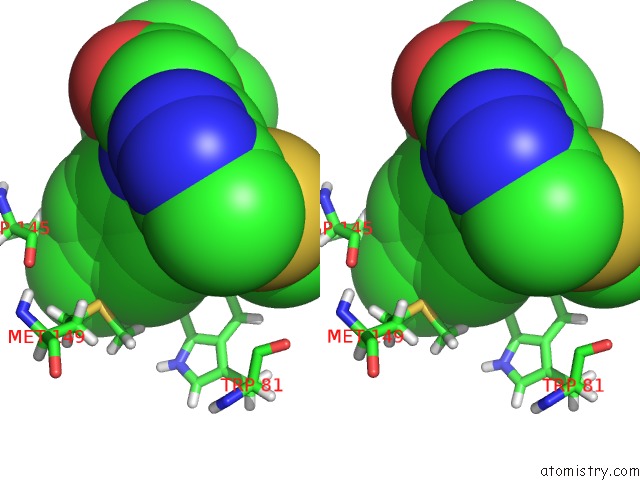
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Crystal Structure of the First Bromodomain of Human 3-Fluoro Tyrosine- Labeled BRD4 in Complex with JQ1 within 5.0Å range:
|
Chlorine binding site 2 out of 2 in 4qzs
Go back to
Chlorine binding site 2 out
of 2 in the Crystal Structure of the First Bromodomain of Human 3-Fluoro Tyrosine- Labeled BRD4 in Complex with JQ1
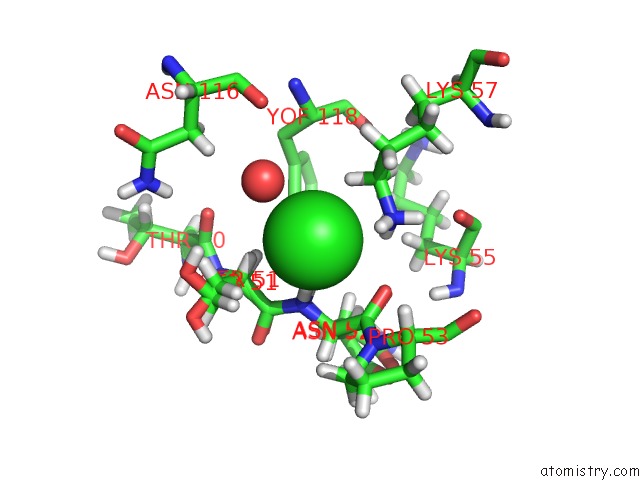
Mono view
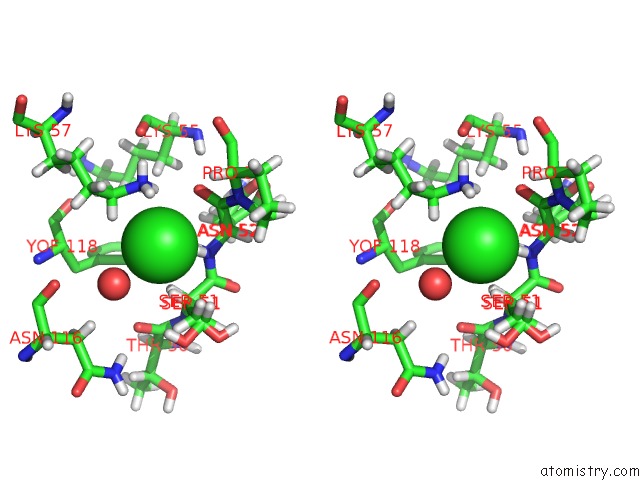
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 2 of Crystal Structure of the First Bromodomain of Human 3-Fluoro Tyrosine- Labeled BRD4 in Complex with JQ1 within 5.0Å range:
|
Reference:
N.K.Mishra,
A.K.Urick,
S.Ember,
E.Schonbrunn,
W.C.Pomerantz.
Fluorinated Aromatic Amino Acids Are Sensitive 19F uc(Nmr) Probes For Bromodomain-Ligand Interactions. Acs Chem.Biol. 2014.
ISSN: ESSN 1554-8937
PubMed: 25290579
DOI: 10.1021/CB5007344
Page generated: Fri Jul 11 21:10:04 2025
ISSN: ESSN 1554-8937
PubMed: 25290579
DOI: 10.1021/CB5007344
Last articles
F in 5BQHF in 5BP0
F in 5B2W
F in 5BOC
F in 5BO9
F in 5B2X
F in 5BNS
F in 5BNR
F in 5BNI
F in 5BMM