Chlorine »
PDB 5ag4-5ao9 »
5am9 »
Chlorine in PDB 5am9: Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16
Enzymatic activity of Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16
All present enzymatic activity of Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16:
3.4.15.1;
3.4.15.1;
Protein crystallography data
The structure of Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16, PDB code: 5am9
was solved by
G.Masuyer,
K.M.Larmuth,
R.G.Douglas,
E.D.Sturrock,
K.R.Acharya,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 113.27 / 1.80 |
| Space group | P 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 73.348, 101.800, 113.950, 85.04, 85.55, 81.88 |
| R / Rfree (%) | 19.674 / 22.907 |
Other elements in 5am9:
The structure of Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16 also contains other interesting chemical elements:
| Zinc | (Zn) | 4 atoms |
| Calcium | (Ca) | 2 atoms |
| Sodium | (Na) | 2 atoms |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16
(pdb code 5am9). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total 4 binding sites of Chlorine where determined in the Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16, PDB code: 5am9:
Jump to Chlorine binding site number: 1; 2; 3; 4;
In total 4 binding sites of Chlorine where determined in the Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16, PDB code: 5am9:
Jump to Chlorine binding site number: 1; 2; 3; 4;
Chlorine binding site 1 out of 4 in 5am9
Go back to
Chlorine binding site 1 out
of 4 in the Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16
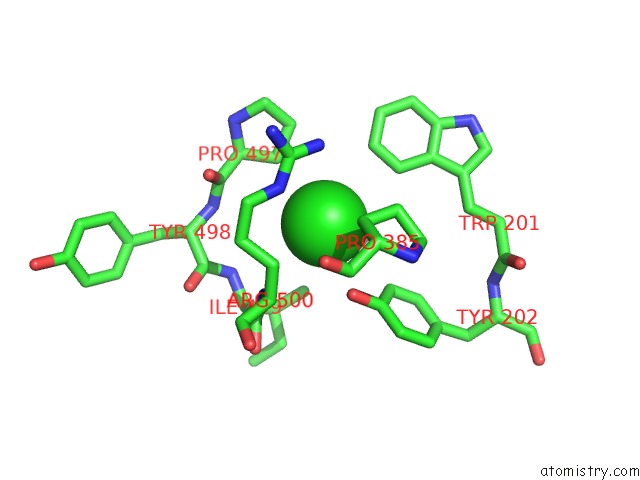
Mono view
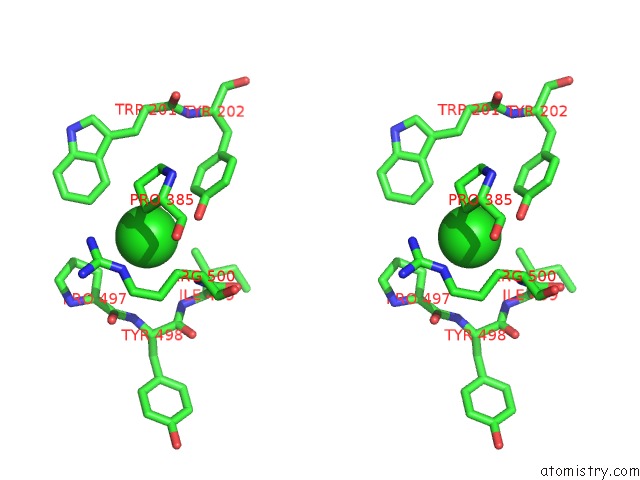
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16 within 5.0Å range:
|
Chlorine binding site 2 out of 4 in 5am9
Go back to
Chlorine binding site 2 out
of 4 in the Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16
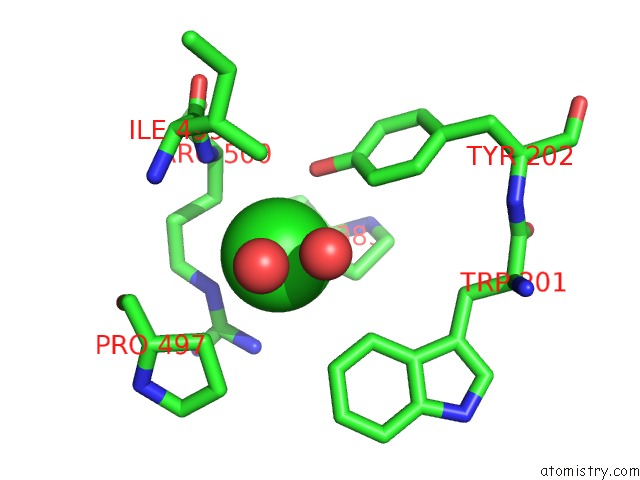
Mono view
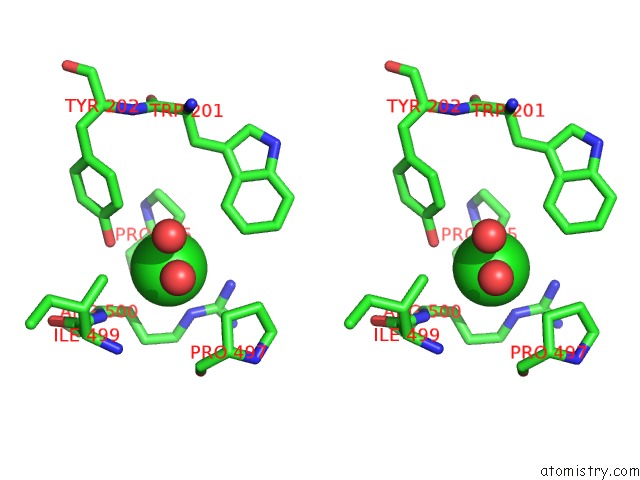
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 2 of Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16 within 5.0Å range:
|
Chlorine binding site 3 out of 4 in 5am9
Go back to
Chlorine binding site 3 out
of 4 in the Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16
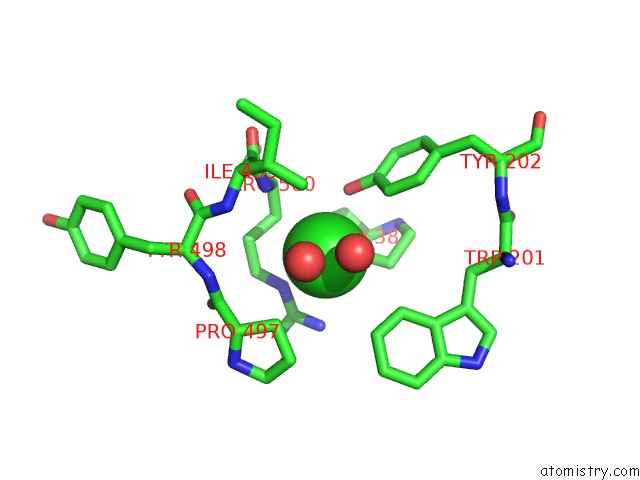
Mono view
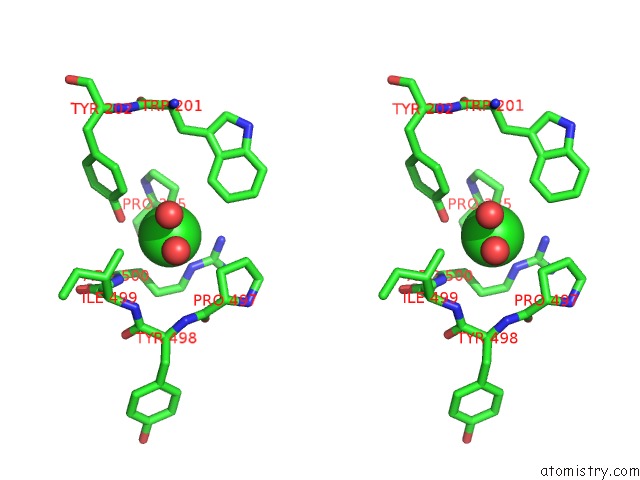
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 3 of Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16 within 5.0Å range:
|
Chlorine binding site 4 out of 4 in 5am9
Go back to
Chlorine binding site 4 out
of 4 in the Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16
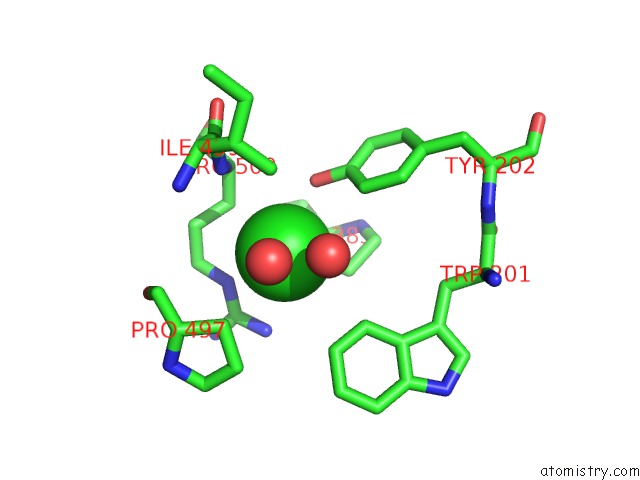
Mono view
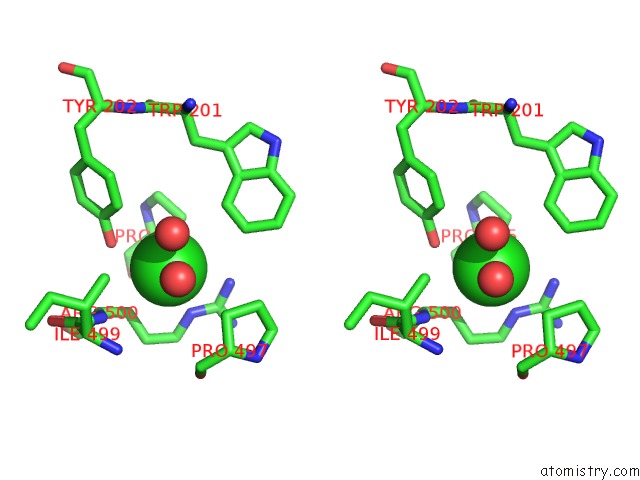
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 4 of Crystal Structure of the Angiotensin-1 Converting Enzyme N- Domain in Complex with Amyloid-Beta 10-16 within 5.0Å range:
|
Reference:
K.M.Larmuth,
G.Masuyer,
R.G.Douglas,
E.D.Sturrock,
K.R.Acharya.
The Kinetic and Structural Characterisation of Amyloid-Beta Metabolism By Human Angiotensin-1- Converting Enzyme (Ace) Febs J. V. 283 1060 2016.
ISSN: ISSN 1742-464X
PubMed: 26748546
DOI: 10.1111/FEBS.13647
Page generated: Sat Jul 12 00:16:02 2025
ISSN: ISSN 1742-464X
PubMed: 26748546
DOI: 10.1111/FEBS.13647
Last articles
F in 7RKWF in 7RKT
F in 7RKR
F in 7RJE
F in 7RJ8
F in 7RH7
F in 7RIV
F in 7RIO
F in 7RIU
F in 7RH0