Chlorine »
PDB 5fzb-5g60 »
5g5g »
Chlorine in PDB 5g5g: Escherichia Coli Periplasmic Aldehyde Oxidase
Protein crystallography data
The structure of Escherichia Coli Periplasmic Aldehyde Oxidase, PDB code: 5g5g
was solved by
M.A.S.Correia,
A.R.Otrelo-Cardoso,
M.J.Romao,
T.Santos-Silva,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 48.32 / 1.70 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 109.681, 78.342, 151.909, 90.00, 99.69, 90.00 |
| R / Rfree (%) | 13.8 / 16.7 |
Other elements in 5g5g:
The structure of Escherichia Coli Periplasmic Aldehyde Oxidase also contains other interesting chemical elements:
| Molybdenum | (Mo) | 1 atom |
| Iodine | (I) | 7 atoms |
| Iron | (Fe) | 8 atoms |
Chlorine Binding Sites:
Pages:
>>> Page 1 <<< Page 2, Binding sites: 11 - 12;Binding sites:
The binding sites of Chlorine atom in the Escherichia Coli Periplasmic Aldehyde Oxidase (pdb code 5g5g). This binding sites where shown within 5.0 Angstroms radius around Chlorine atom.In total 12 binding sites of Chlorine where determined in the Escherichia Coli Periplasmic Aldehyde Oxidase, PDB code: 5g5g:
Jump to Chlorine binding site number: 1; 2; 3; 4; 5; 6; 7; 8; 9; 10;
Chlorine binding site 1 out of 12 in 5g5g
Go back to
Chlorine binding site 1 out
of 12 in the Escherichia Coli Periplasmic Aldehyde Oxidase
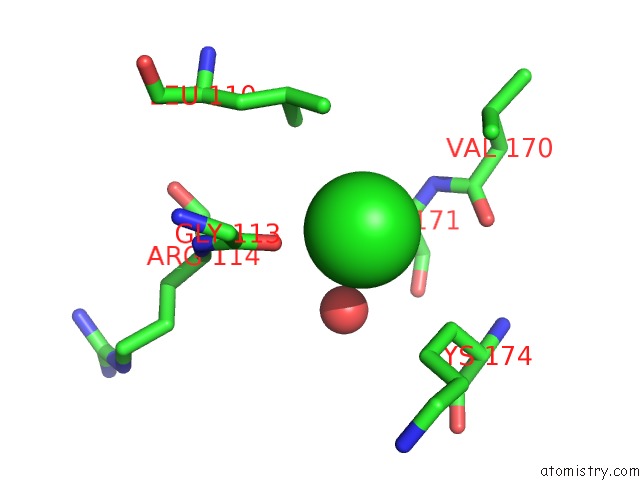
Mono view
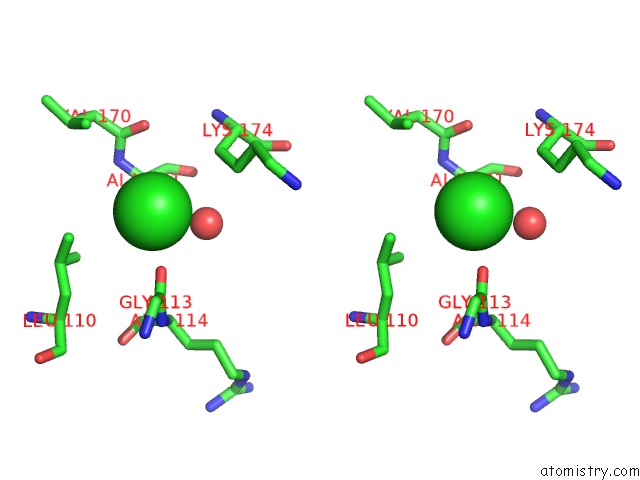
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Escherichia Coli Periplasmic Aldehyde Oxidase within 5.0Å range:
|
Chlorine binding site 2 out of 12 in 5g5g
Go back to
Chlorine binding site 2 out
of 12 in the Escherichia Coli Periplasmic Aldehyde Oxidase
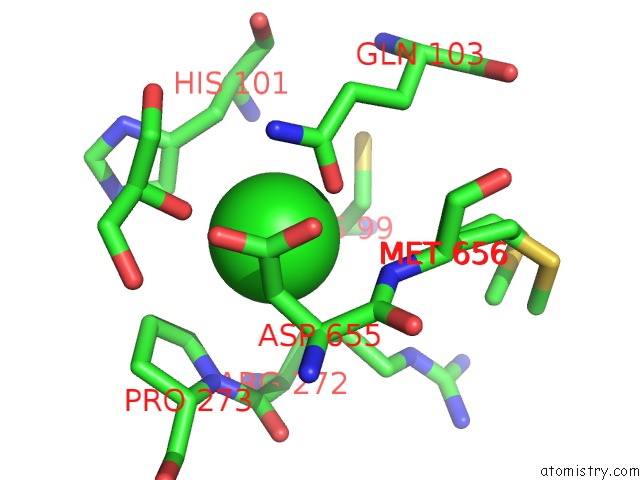
Mono view
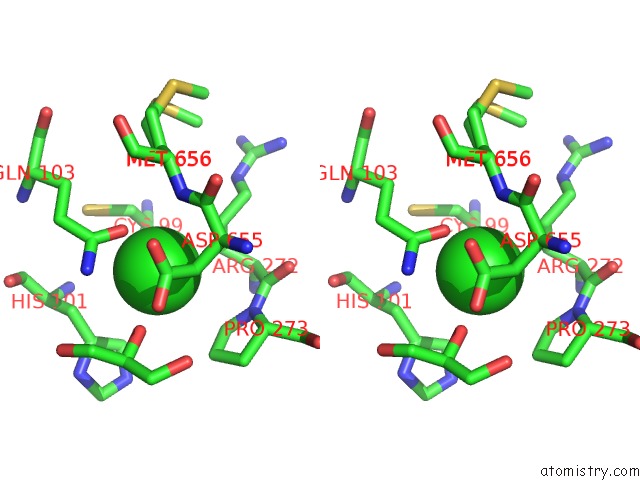
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 2 of Escherichia Coli Periplasmic Aldehyde Oxidase within 5.0Å range:
|
Chlorine binding site 3 out of 12 in 5g5g
Go back to
Chlorine binding site 3 out
of 12 in the Escherichia Coli Periplasmic Aldehyde Oxidase
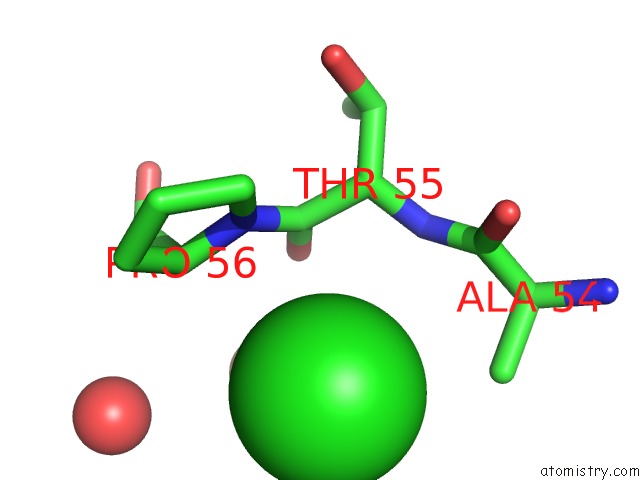
Mono view
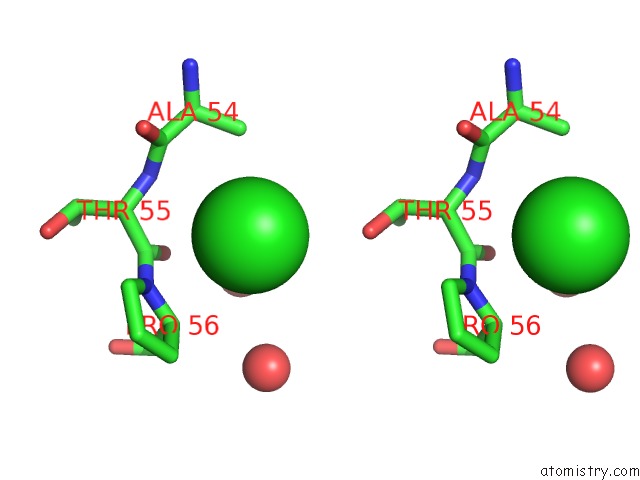
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 3 of Escherichia Coli Periplasmic Aldehyde Oxidase within 5.0Å range:
|
Chlorine binding site 4 out of 12 in 5g5g
Go back to
Chlorine binding site 4 out
of 12 in the Escherichia Coli Periplasmic Aldehyde Oxidase
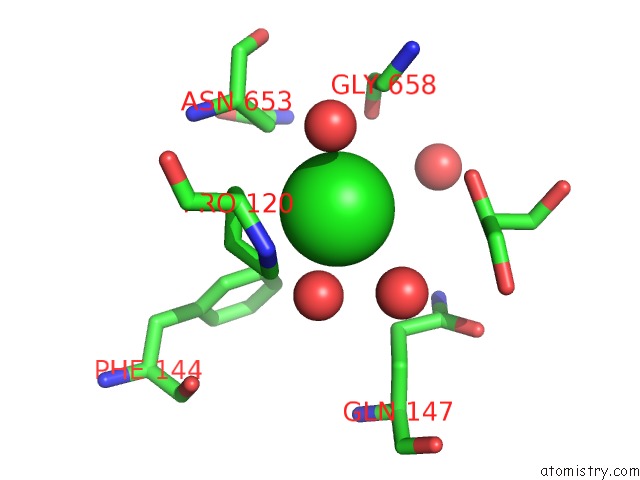
Mono view
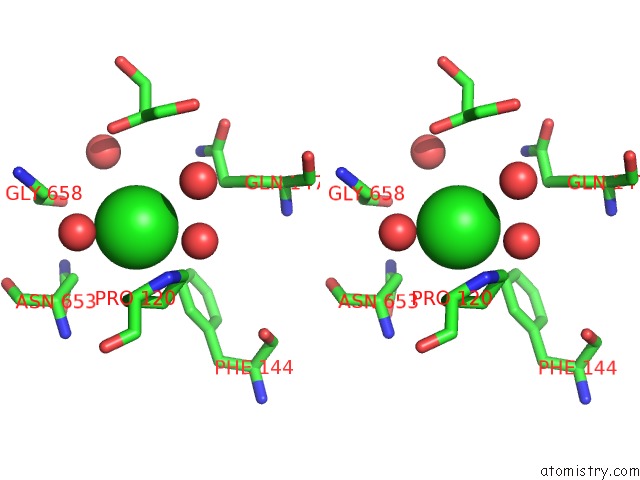
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 4 of Escherichia Coli Periplasmic Aldehyde Oxidase within 5.0Å range:
|
Chlorine binding site 5 out of 12 in 5g5g
Go back to
Chlorine binding site 5 out
of 12 in the Escherichia Coli Periplasmic Aldehyde Oxidase
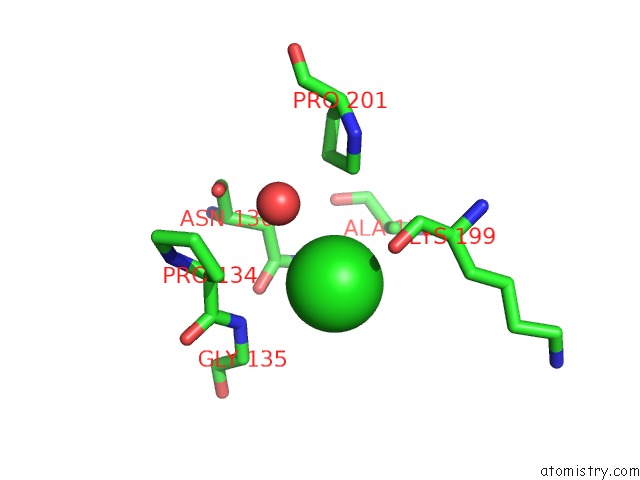
Mono view
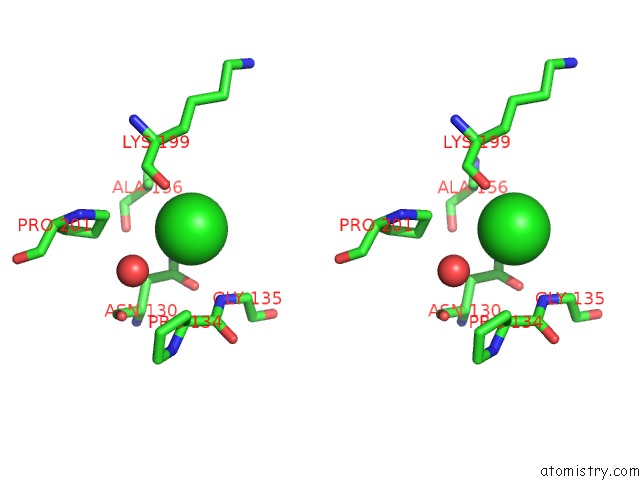
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 5 of Escherichia Coli Periplasmic Aldehyde Oxidase within 5.0Å range:
|
Chlorine binding site 6 out of 12 in 5g5g
Go back to
Chlorine binding site 6 out
of 12 in the Escherichia Coli Periplasmic Aldehyde Oxidase
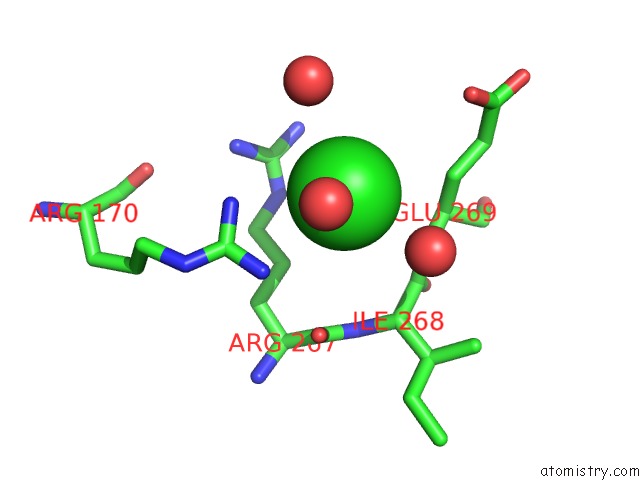
Mono view
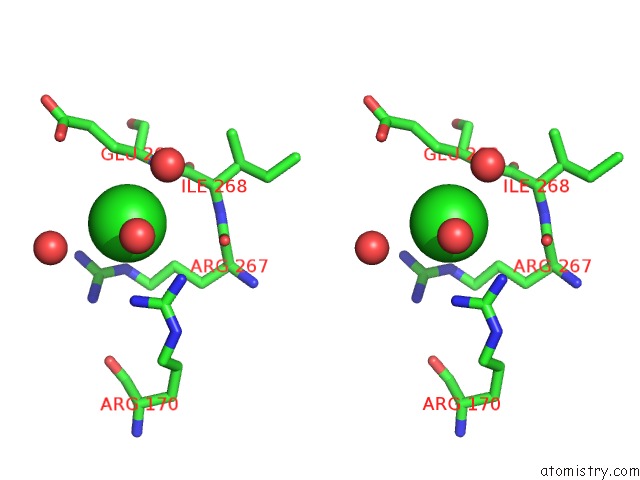
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 6 of Escherichia Coli Periplasmic Aldehyde Oxidase within 5.0Å range:
|
Chlorine binding site 7 out of 12 in 5g5g
Go back to
Chlorine binding site 7 out
of 12 in the Escherichia Coli Periplasmic Aldehyde Oxidase
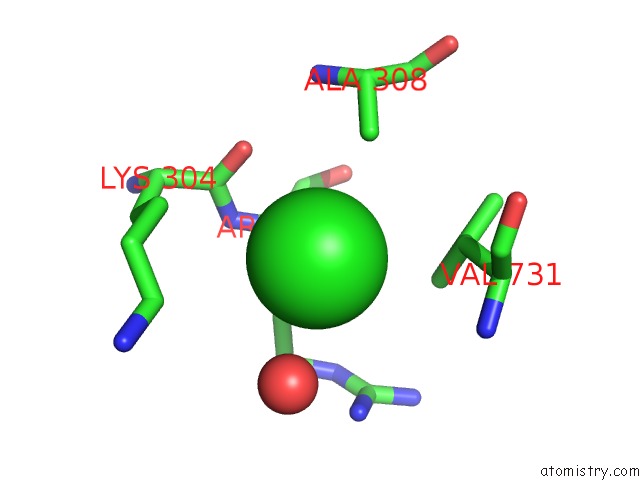
Mono view
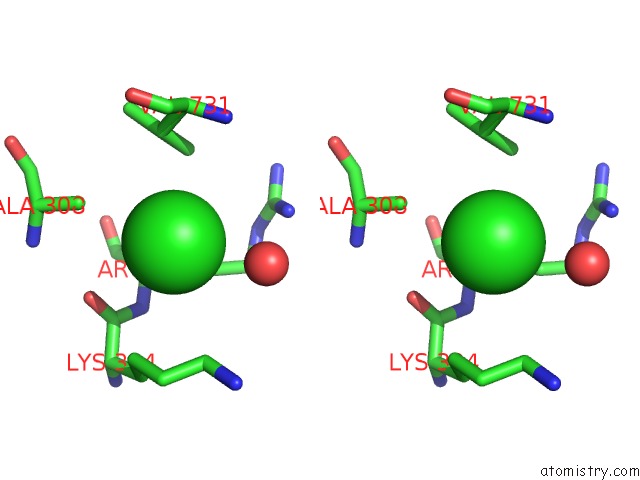
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 7 of Escherichia Coli Periplasmic Aldehyde Oxidase within 5.0Å range:
|
Chlorine binding site 8 out of 12 in 5g5g
Go back to
Chlorine binding site 8 out
of 12 in the Escherichia Coli Periplasmic Aldehyde Oxidase
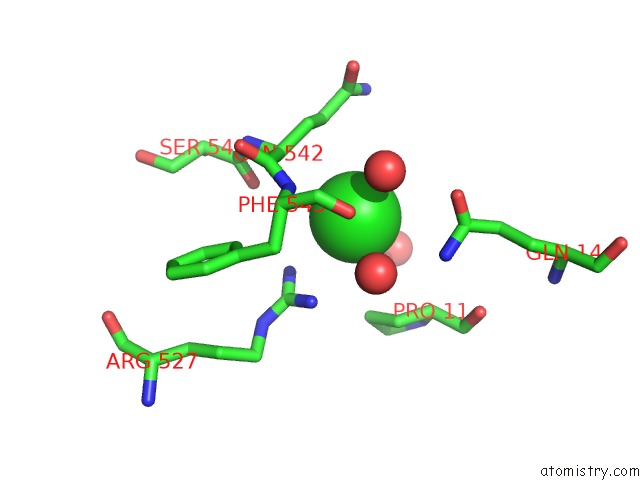
Mono view
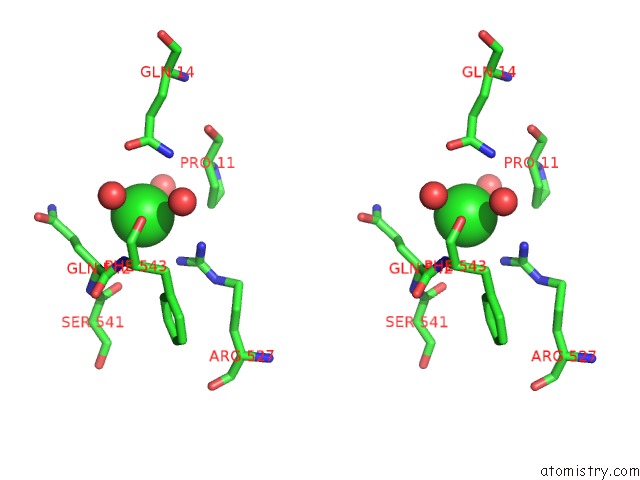
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 8 of Escherichia Coli Periplasmic Aldehyde Oxidase within 5.0Å range:
|
Chlorine binding site 9 out of 12 in 5g5g
Go back to
Chlorine binding site 9 out
of 12 in the Escherichia Coli Periplasmic Aldehyde Oxidase
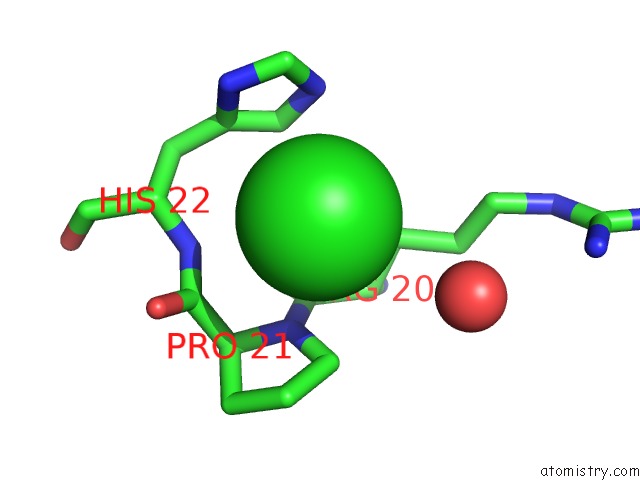
Mono view
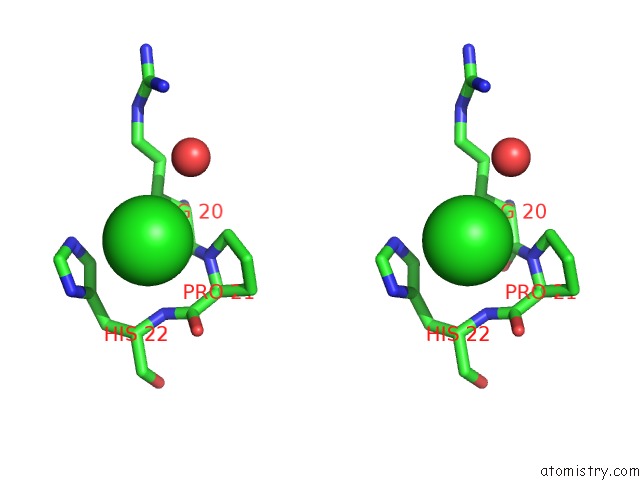
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 9 of Escherichia Coli Periplasmic Aldehyde Oxidase within 5.0Å range:
|
Chlorine binding site 10 out of 12 in 5g5g
Go back to
Chlorine binding site 10 out
of 12 in the Escherichia Coli Periplasmic Aldehyde Oxidase
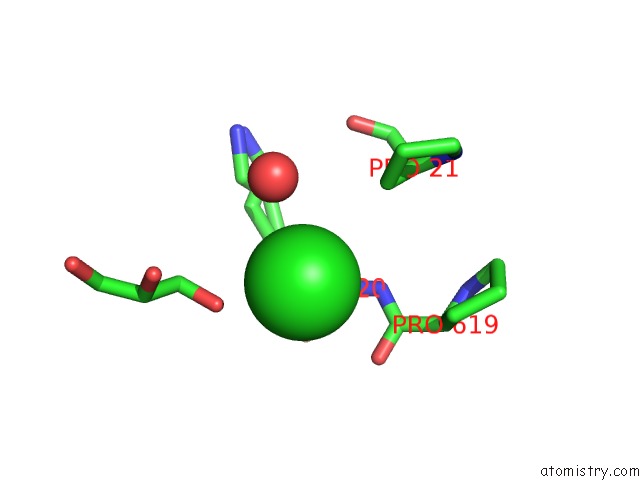
Mono view
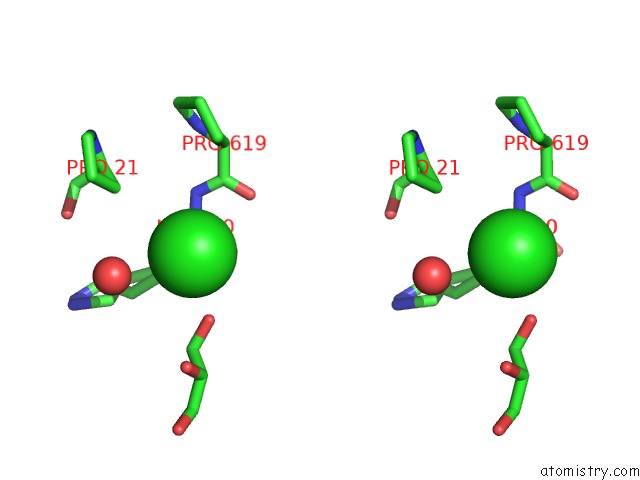
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 10 of Escherichia Coli Periplasmic Aldehyde Oxidase within 5.0Å range:
|
Reference:
M.A.Correia,
A.R.Otrelo-Cardoso,
V.Schwuchow,
K.G.Sigfridsson Clauss,
M.Haumann,
M.J.Romao,
S.Leimkuhler,
T.Santos-Silva.
The Escherichia Coli Periplasmic Aldehyde Oxidoreductase Is An Exceptional Member of the Xanthine Oxidase Family of Molybdoenzymes. Acs Chem.Biol. V. 11 2923 2016.
ISSN: ISSN 1554-8929
PubMed: 27622978
DOI: 10.1021/ACSCHEMBIO.6B00572
Page generated: Fri Jul 26 08:20:48 2024
ISSN: ISSN 1554-8929
PubMed: 27622978
DOI: 10.1021/ACSCHEMBIO.6B00572
Last articles
Ca in 5MWCCa in 5MWB
Ca in 5MWK
Ca in 5MW5
Ca in 5MUW
Ca in 5MVV
Ca in 5MUV
Ca in 5MUA
Ca in 5MSX
Ca in 5MU9