Chlorine »
PDB 5xii-5xvf »
5xio »
Chlorine in PDB 5xio: Crystal Structure of Cryptosporidium Parvum Prolyl-Trna Synthetase (Cpprs) in Complex with Halofuginone
Protein crystallography data
The structure of Crystal Structure of Cryptosporidium Parvum Prolyl-Trna Synthetase (Cpprs) in Complex with Halofuginone, PDB code: 5xio
was solved by
V.Jain,
Y.Manickam,
A.Sharma,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 29.31 / 2.46 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 82.768, 112.635, 140.494, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 19.6 / 25.2 |
Other elements in 5xio:
The structure of Crystal Structure of Cryptosporidium Parvum Prolyl-Trna Synthetase (Cpprs) in Complex with Halofuginone also contains other interesting chemical elements:
| Zinc | (Zn) | 2 atoms |
| Bromine | (Br) | 2 atoms |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Crystal Structure of Cryptosporidium Parvum Prolyl-Trna Synthetase (Cpprs) in Complex with Halofuginone
(pdb code 5xio). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total 2 binding sites of Chlorine where determined in the Crystal Structure of Cryptosporidium Parvum Prolyl-Trna Synthetase (Cpprs) in Complex with Halofuginone, PDB code: 5xio:
Jump to Chlorine binding site number: 1; 2;
In total 2 binding sites of Chlorine where determined in the Crystal Structure of Cryptosporidium Parvum Prolyl-Trna Synthetase (Cpprs) in Complex with Halofuginone, PDB code: 5xio:
Jump to Chlorine binding site number: 1; 2;
Chlorine binding site 1 out of 2 in 5xio
Go back to
Chlorine binding site 1 out
of 2 in the Crystal Structure of Cryptosporidium Parvum Prolyl-Trna Synthetase (Cpprs) in Complex with Halofuginone
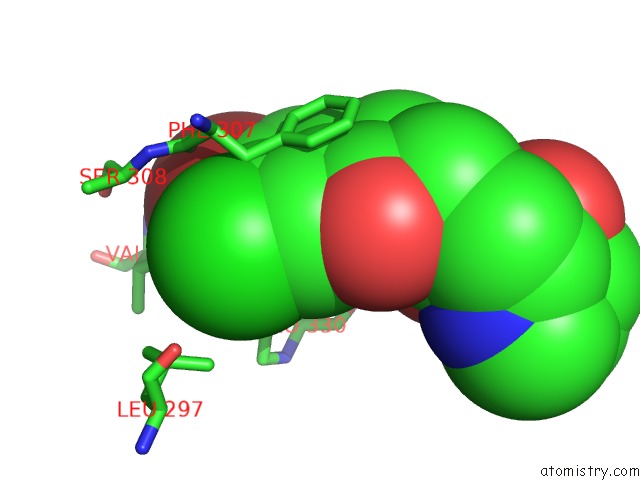
Mono view
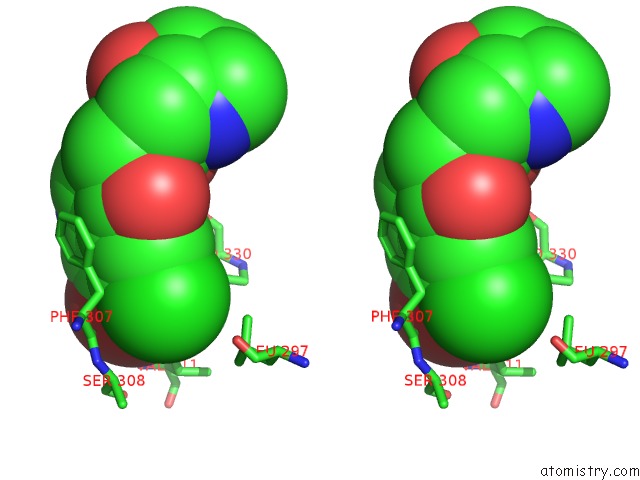
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Crystal Structure of Cryptosporidium Parvum Prolyl-Trna Synthetase (Cpprs) in Complex with Halofuginone within 5.0Å range:
|
Chlorine binding site 2 out of 2 in 5xio
Go back to
Chlorine binding site 2 out
of 2 in the Crystal Structure of Cryptosporidium Parvum Prolyl-Trna Synthetase (Cpprs) in Complex with Halofuginone
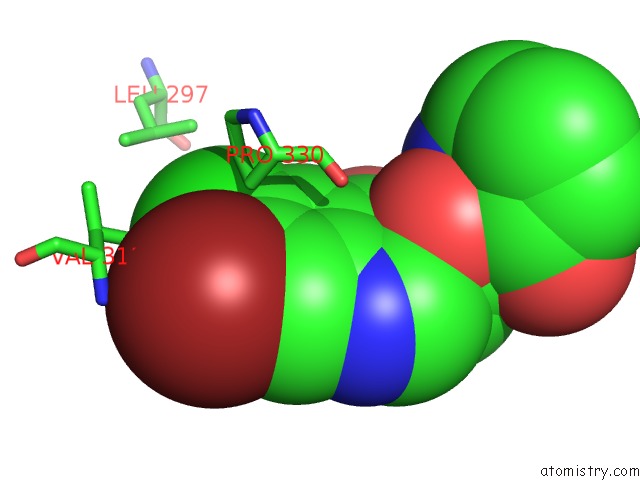
Mono view
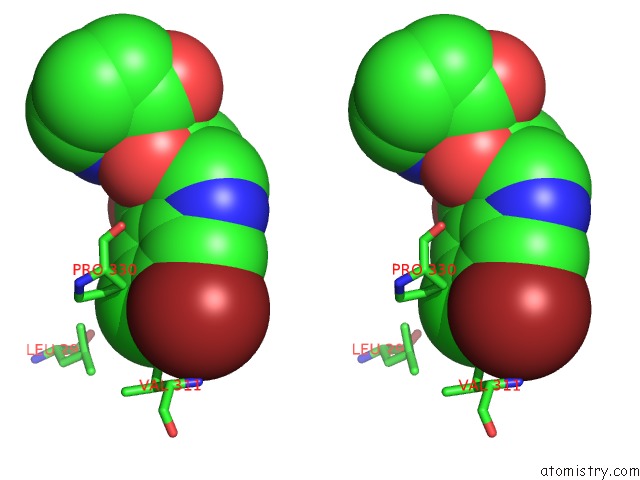
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 2 of Crystal Structure of Cryptosporidium Parvum Prolyl-Trna Synthetase (Cpprs) in Complex with Halofuginone within 5.0Å range:
|
Reference:
V.Jain,
M.Yogavel,
H.Kikuchi,
Y.Oshima,
N.Hariguchi,
M.Matsumoto,
P.Goel,
B.Touquet,
R.S.Jumani,
F.Tacchini-Cottier,
K.Harlos,
C.D.Huston,
M.A.Hakimi,
A.Sharma.
Targeting Prolyl-Trna Synthetase to Accelerate Drug Discovery Against Malaria, Leishmaniasis, Toxoplasmosis, Cryptosporidiosis, and Coccidiosis Structure V. 25 1495 2017.
ISSN: ISSN 1878-4186
PubMed: 28867614
DOI: 10.1016/J.STR.2017.07.015
Page generated: Sat Jul 12 10:40:19 2025
ISSN: ISSN 1878-4186
PubMed: 28867614
DOI: 10.1016/J.STR.2017.07.015
Last articles
F in 4IFVF in 4IDO
F in 4ICC
F in 4IBI
F in 4IAH
F in 4IAE
F in 4I9H
F in 4I9N
F in 4I9O
F in 4IA9