Chlorine »
PDB 6de4-6dov »
6dif »
Chlorine in PDB 6dif: Wild-Type Hiv-1 Protease in Complex with Tipranavir
Protein crystallography data
The structure of Wild-Type Hiv-1 Protease in Complex with Tipranavir, PDB code: 6dif
was solved by
A.E.Wong-Sam,
Y.-F.Wang,
I.T.Weber,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 25.00 / 1.20 |
| Space group | P 21 21 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 58.300, 86.259, 46.005, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 15.4 / 18.7 |
Other elements in 6dif:
The structure of Wild-Type Hiv-1 Protease in Complex with Tipranavir also contains other interesting chemical elements:
| Fluorine | (F) | 6 atoms |
| Sodium | (Na) | 1 atom |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Wild-Type Hiv-1 Protease in Complex with Tipranavir
(pdb code 6dif). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total 3 binding sites of Chlorine where determined in the Wild-Type Hiv-1 Protease in Complex with Tipranavir, PDB code: 6dif:
Jump to Chlorine binding site number: 1; 2; 3;
In total 3 binding sites of Chlorine where determined in the Wild-Type Hiv-1 Protease in Complex with Tipranavir, PDB code: 6dif:
Jump to Chlorine binding site number: 1; 2; 3;
Chlorine binding site 1 out of 3 in 6dif
Go back to
Chlorine binding site 1 out
of 3 in the Wild-Type Hiv-1 Protease in Complex with Tipranavir
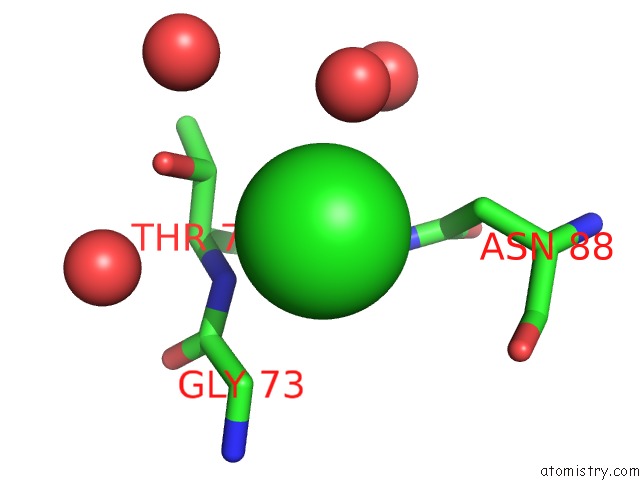
Mono view
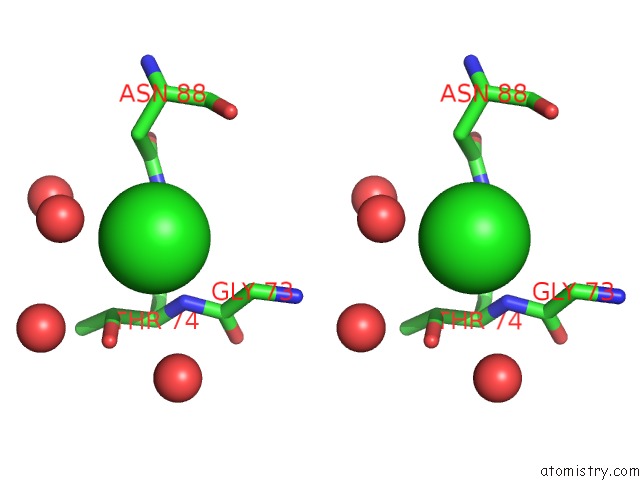
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Wild-Type Hiv-1 Protease in Complex with Tipranavir within 5.0Å range:
|
Chlorine binding site 2 out of 3 in 6dif
Go back to
Chlorine binding site 2 out
of 3 in the Wild-Type Hiv-1 Protease in Complex with Tipranavir
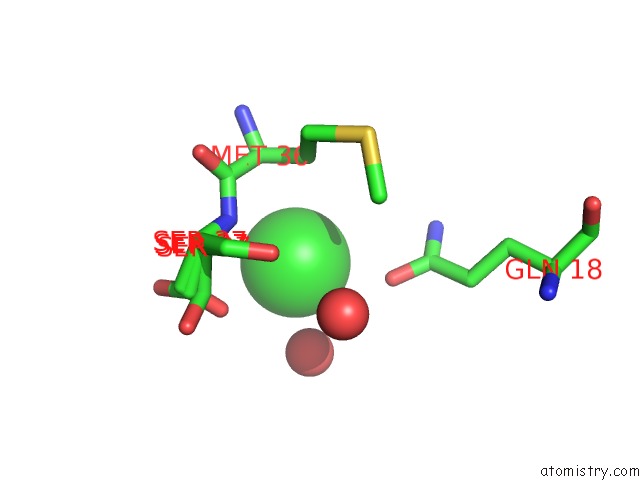
Mono view
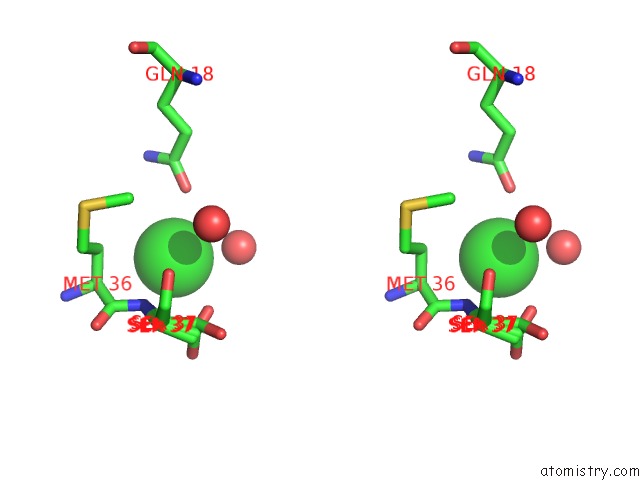
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 2 of Wild-Type Hiv-1 Protease in Complex with Tipranavir within 5.0Å range:
|
Chlorine binding site 3 out of 3 in 6dif
Go back to
Chlorine binding site 3 out
of 3 in the Wild-Type Hiv-1 Protease in Complex with Tipranavir
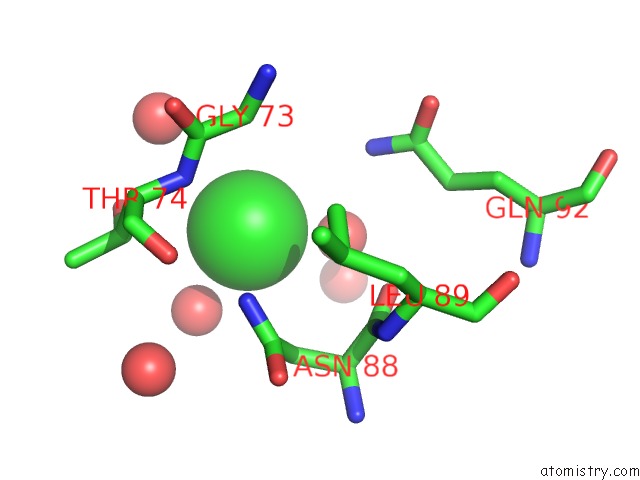
Mono view
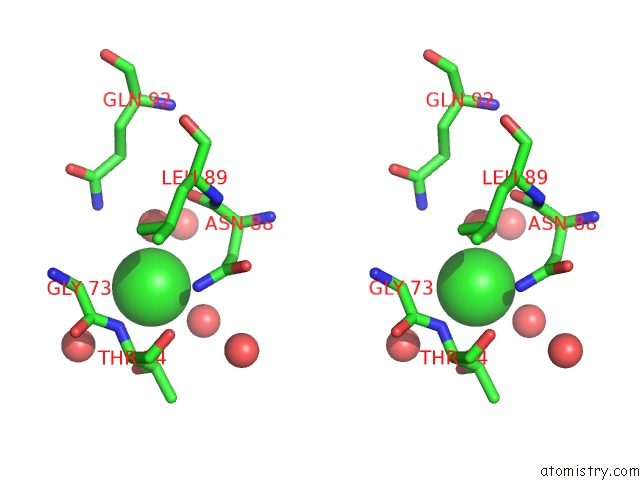
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 3 of Wild-Type Hiv-1 Protease in Complex with Tipranavir within 5.0Å range:
|
Reference:
A.Wong-Sam,
Y.F.Wang,
Y.Zhang,
A.K.Ghosh,
R.W.Harrison,
I.T.Weber.
Drug Resistance Mutation L76V Alters Nonpolar Interactions at the Flap-Core Interface of Hiv-1 Protease. Acs Omega V. 3 12132 2018.
ISSN: ESSN 2470-1343
PubMed: 30288468
DOI: 10.1021/ACSOMEGA.8B01683
Page generated: Sat Jul 27 21:32:26 2024
ISSN: ESSN 2470-1343
PubMed: 30288468
DOI: 10.1021/ACSOMEGA.8B01683
Last articles
Ca in 5MFACa in 5MF4
Ca in 5MEY
Ca in 5MB1
Ca in 5MAZ
Ca in 5MER
Ca in 5MAY
Ca in 5MEH
Ca in 5MC9
Ca in 5MA7