Chlorine »
PDB 6h4j-6hb2 »
6h9a »
Chlorine in PDB 6h9a: Crystal Structure of Anaerobic Ergothioneine Biosynthesis Enzyme From Chlorobium Limicola in Complex with Natural Substrate Trimethyl Histidine.
Protein crystallography data
The structure of Crystal Structure of Anaerobic Ergothioneine Biosynthesis Enzyme From Chlorobium Limicola in Complex with Natural Substrate Trimethyl Histidine., PDB code: 6h9a
was solved by
F.Leisinger,
R.Burn,
M.Meury,
P.Lukat,
F.P.Seebeck,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 46.94 / 2.83 |
| Space group | P 21 21 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 76.550, 178.240, 43.266, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 18.7 / 21.7 |
Other elements in 6h9a:
The structure of Crystal Structure of Anaerobic Ergothioneine Biosynthesis Enzyme From Chlorobium Limicola in Complex with Natural Substrate Trimethyl Histidine. also contains other interesting chemical elements:
| Sodium | (Na) | 1 atom |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Crystal Structure of Anaerobic Ergothioneine Biosynthesis Enzyme From Chlorobium Limicola in Complex with Natural Substrate Trimethyl Histidine.
(pdb code 6h9a). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total 3 binding sites of Chlorine where determined in the Crystal Structure of Anaerobic Ergothioneine Biosynthesis Enzyme From Chlorobium Limicola in Complex with Natural Substrate Trimethyl Histidine., PDB code: 6h9a:
Jump to Chlorine binding site number: 1; 2; 3;
In total 3 binding sites of Chlorine where determined in the Crystal Structure of Anaerobic Ergothioneine Biosynthesis Enzyme From Chlorobium Limicola in Complex with Natural Substrate Trimethyl Histidine., PDB code: 6h9a:
Jump to Chlorine binding site number: 1; 2; 3;
Chlorine binding site 1 out of 3 in 6h9a
Go back to
Chlorine binding site 1 out
of 3 in the Crystal Structure of Anaerobic Ergothioneine Biosynthesis Enzyme From Chlorobium Limicola in Complex with Natural Substrate Trimethyl Histidine.
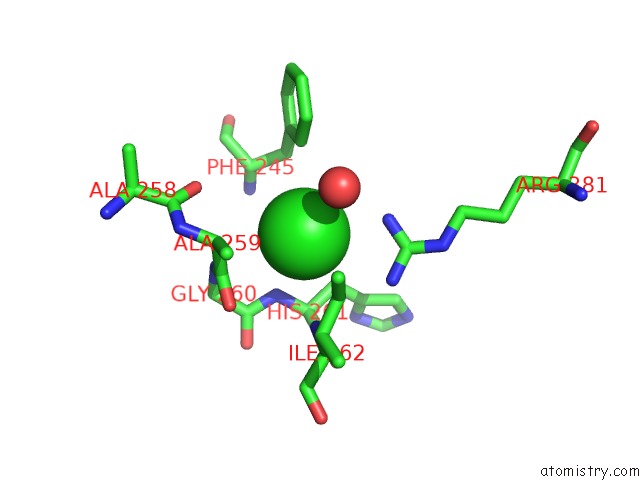
Mono view
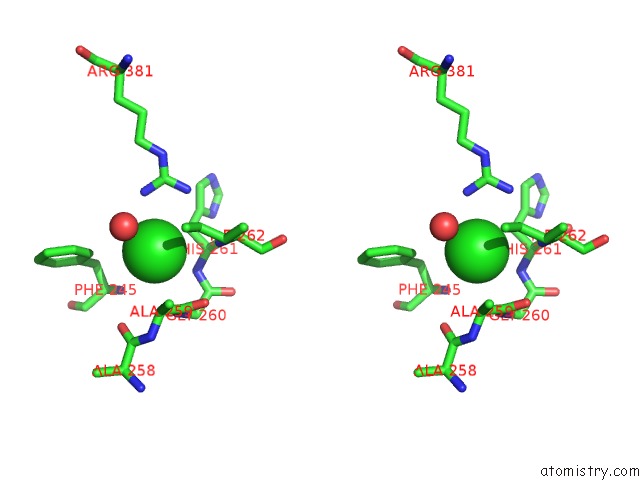
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Crystal Structure of Anaerobic Ergothioneine Biosynthesis Enzyme From Chlorobium Limicola in Complex with Natural Substrate Trimethyl Histidine. within 5.0Å range:
|
Chlorine binding site 2 out of 3 in 6h9a
Go back to
Chlorine binding site 2 out
of 3 in the Crystal Structure of Anaerobic Ergothioneine Biosynthesis Enzyme From Chlorobium Limicola in Complex with Natural Substrate Trimethyl Histidine.
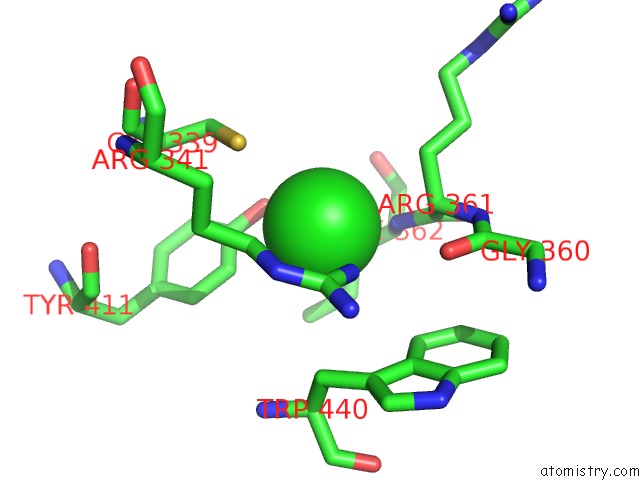
Mono view
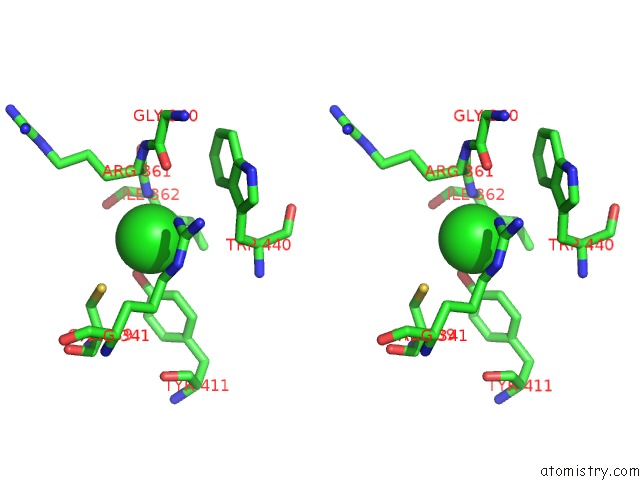
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 2 of Crystal Structure of Anaerobic Ergothioneine Biosynthesis Enzyme From Chlorobium Limicola in Complex with Natural Substrate Trimethyl Histidine. within 5.0Å range:
|
Chlorine binding site 3 out of 3 in 6h9a
Go back to
Chlorine binding site 3 out
of 3 in the Crystal Structure of Anaerobic Ergothioneine Biosynthesis Enzyme From Chlorobium Limicola in Complex with Natural Substrate Trimethyl Histidine.
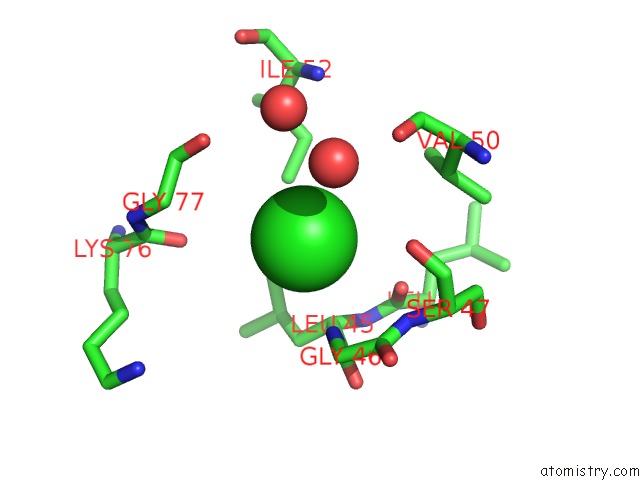
Mono view
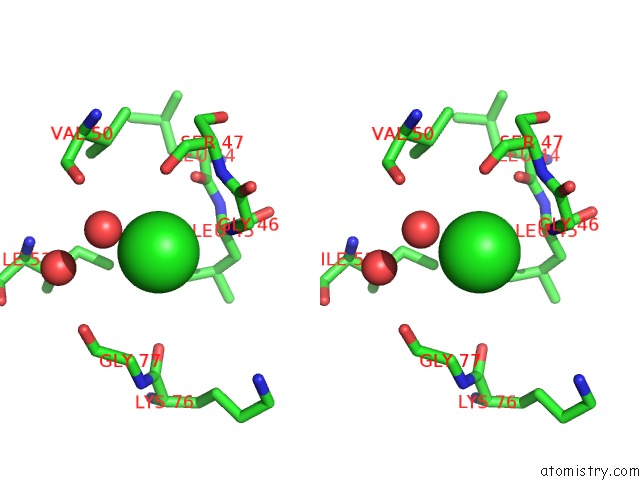
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 3 of Crystal Structure of Anaerobic Ergothioneine Biosynthesis Enzyme From Chlorobium Limicola in Complex with Natural Substrate Trimethyl Histidine. within 5.0Å range:
|
Reference:
F.Leisinger,
R.Burn,
M.Meury,
P.Lukat,
F.P.Seebeck.
Structural and Mechanistic Basis For Anaerobic Ergothioneine Biosynthesis. J.Am.Chem.Soc. V. 141 6906 2019.
ISSN: ESSN 1520-5126
PubMed: 30943021
DOI: 10.1021/JACS.8B12596
Page generated: Sun Jul 28 00:42:03 2024
ISSN: ESSN 1520-5126
PubMed: 30943021
DOI: 10.1021/JACS.8B12596
Last articles
Cl in 2WGSCl in 2WIU
Cl in 2WJP
Cl in 2WJ2
Cl in 2WJ9
Cl in 2WJ1
Cl in 2WIK
Cl in 2WIL
Cl in 2WIG
Cl in 2WIJ