Chlorine »
PDB 6m7z-6miy »
6mgj »
Chlorine in PDB 6mgj: Crystal Structure of the Catalytic Domain From GH74 Enzyme POGH74 From Paenibacillus Odorifer, Apoenzyme
Protein crystallography data
The structure of Crystal Structure of the Catalytic Domain From GH74 Enzyme POGH74 From Paenibacillus Odorifer, Apoenzyme, PDB code: 6mgj
was solved by
P.J.Stogios,
T.Skarina,
B.Nocek,
G.Arnal,
H.Brumer,
A.Savchenko,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 39.78 / 2.00 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 239.094, 226.674, 187.224, 90.00, 114.07, 90.00 |
| R / Rfree (%) | 14.9 / 17.7 |
Other elements in 6mgj:
The structure of Crystal Structure of the Catalytic Domain From GH74 Enzyme POGH74 From Paenibacillus Odorifer, Apoenzyme also contains other interesting chemical elements:
| Magnesium | (Mg) | 9 atoms |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Crystal Structure of the Catalytic Domain From GH74 Enzyme POGH74 From Paenibacillus Odorifer, Apoenzyme
(pdb code 6mgj). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total only one binding site of Chlorine was determined in the Crystal Structure of the Catalytic Domain From GH74 Enzyme POGH74 From Paenibacillus Odorifer, Apoenzyme, PDB code: 6mgj:
In total only one binding site of Chlorine was determined in the Crystal Structure of the Catalytic Domain From GH74 Enzyme POGH74 From Paenibacillus Odorifer, Apoenzyme, PDB code: 6mgj:
Chlorine binding site 1 out of 1 in 6mgj
Go back to
Chlorine binding site 1 out
of 1 in the Crystal Structure of the Catalytic Domain From GH74 Enzyme POGH74 From Paenibacillus Odorifer, Apoenzyme

Mono view
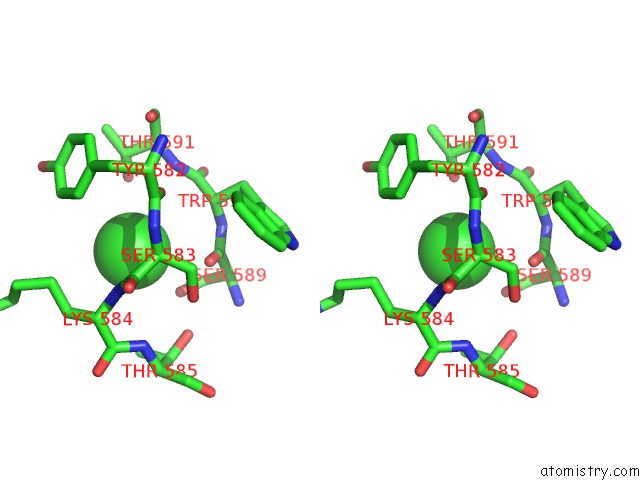
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Crystal Structure of the Catalytic Domain From GH74 Enzyme POGH74 From Paenibacillus Odorifer, Apoenzyme within 5.0Å range:
|
Reference:
G.Arnal,
P.J.Stogios,
J.Asohan,
T.Skarina,
A.Savchenko,
H.Brumer.
Structural Enzymology Reveals the Molecular Basis of Substrate Regiospecificity and Processivity of An Exemplar Bacterial Glycoside Hydrolase Family 74ENDO-Xyloglucanase. Biochem. J. V. 475 3963 2018.
ISSN: ESSN 1470-8728
PubMed: 30463871
DOI: 10.1042/BCJ20180763
Page generated: Sun Jul 28 03:12:48 2024
ISSN: ESSN 1470-8728
PubMed: 30463871
DOI: 10.1042/BCJ20180763
Last articles
Zn in 9JYWZn in 9IR4
Zn in 9IR3
Zn in 9GMX
Zn in 9GMW
Zn in 9JEJ
Zn in 9ERF
Zn in 9ERE
Zn in 9EGV
Zn in 9EGW