Chlorine »
PDB 8fwn-8ga3 »
8fww »
Chlorine in PDB 8fww: Structure of the Ligand-Binding and Transmembrane Domains of Kainate Receptor GLUK2 in Complex with the Positive Allosteric Modulator BPAM344 and Noncompetitive Inhibitor Perampanel
Other elements in 8fww:
The structure of Structure of the Ligand-Binding and Transmembrane Domains of Kainate Receptor GLUK2 in Complex with the Positive Allosteric Modulator BPAM344 and Noncompetitive Inhibitor Perampanel also contains other interesting chemical elements:
| Sodium | (Na) | 7 atoms |
| Fluorine | (F) | 4 atoms |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Structure of the Ligand-Binding and Transmembrane Domains of Kainate Receptor GLUK2 in Complex with the Positive Allosteric Modulator BPAM344 and Noncompetitive Inhibitor Perampanel
(pdb code 8fww). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total 2 binding sites of Chlorine where determined in the Structure of the Ligand-Binding and Transmembrane Domains of Kainate Receptor GLUK2 in Complex with the Positive Allosteric Modulator BPAM344 and Noncompetitive Inhibitor Perampanel, PDB code: 8fww:
Jump to Chlorine binding site number: 1; 2;
In total 2 binding sites of Chlorine where determined in the Structure of the Ligand-Binding and Transmembrane Domains of Kainate Receptor GLUK2 in Complex with the Positive Allosteric Modulator BPAM344 and Noncompetitive Inhibitor Perampanel, PDB code: 8fww:
Jump to Chlorine binding site number: 1; 2;
Chlorine binding site 1 out of 2 in 8fww
Go back to
Chlorine binding site 1 out
of 2 in the Structure of the Ligand-Binding and Transmembrane Domains of Kainate Receptor GLUK2 in Complex with the Positive Allosteric Modulator BPAM344 and Noncompetitive Inhibitor Perampanel

Mono view
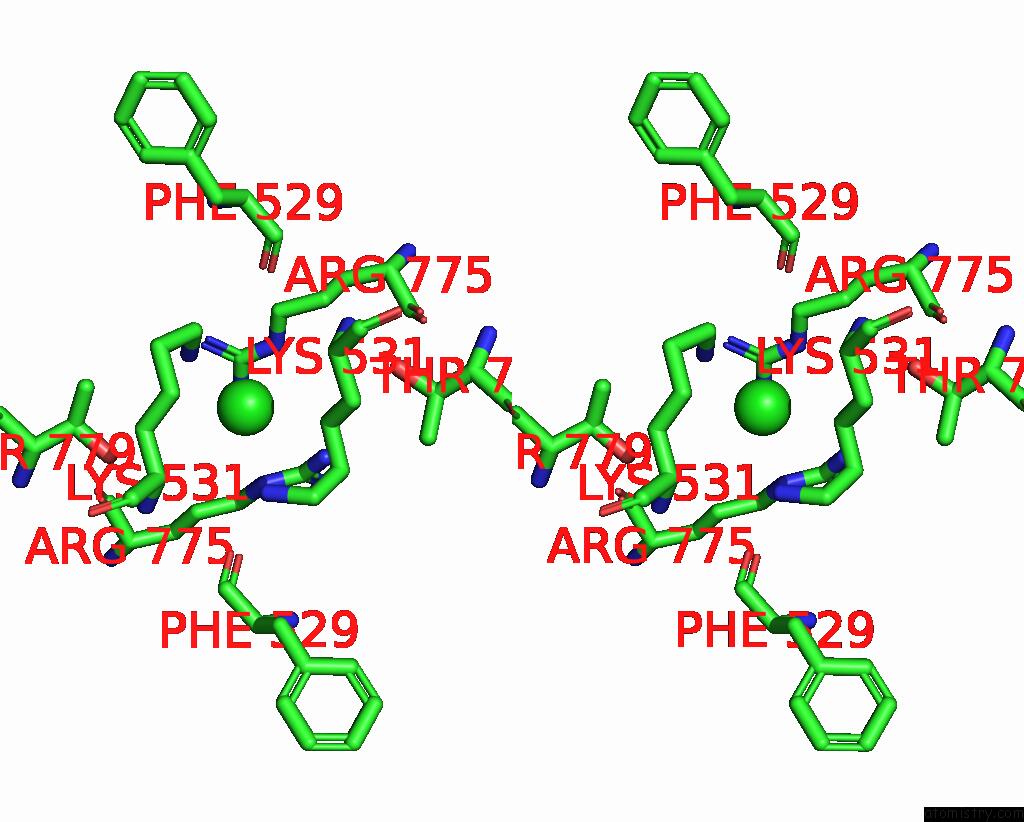
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Structure of the Ligand-Binding and Transmembrane Domains of Kainate Receptor GLUK2 in Complex with the Positive Allosteric Modulator BPAM344 and Noncompetitive Inhibitor Perampanel within 5.0Å range:
|
Chlorine binding site 2 out of 2 in 8fww
Go back to
Chlorine binding site 2 out
of 2 in the Structure of the Ligand-Binding and Transmembrane Domains of Kainate Receptor GLUK2 in Complex with the Positive Allosteric Modulator BPAM344 and Noncompetitive Inhibitor Perampanel

Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 2 of Structure of the Ligand-Binding and Transmembrane Domains of Kainate Receptor GLUK2 in Complex with the Positive Allosteric Modulator BPAM344 and Noncompetitive Inhibitor Perampanel within 5.0Å range:
|
Reference:
S.P.Gangwar,
L.Y.Yen,
M.V.Yelshanskaya,
A.I.Sobolevsky.
Positive and Negative Allosteric Modulation of GLUK2 Kainate Receptors By BPAM344 and Antiepileptic Perampanel. Cell Rep V. 42 12124 2023.
ISSN: ESSN 2211-1247
PubMed: 36857176
DOI: 10.1016/J.CELREP.2023.112124
Page generated: Sun Jul 13 11:41:59 2025
ISSN: ESSN 2211-1247
PubMed: 36857176
DOI: 10.1016/J.CELREP.2023.112124
Last articles
F in 7Q4AF in 7PV7
F in 7Q2Y
F in 7Q3B
F in 7Q2X
F in 7PVK
F in 7Q2J
F in 7Q01
F in 7PZX
F in 7PZW