Chlorine »
PDB 8wtq-8xar »
8x64 »
Chlorine in PDB 8x64: Cryoem Structure of the Histamine H1 Receptor-Bril/Anti Bril Fab Complex with Desloratadine
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Cryoem Structure of the Histamine H1 Receptor-Bril/Anti Bril Fab Complex with Desloratadine
(pdb code 8x64). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total only one binding site of Chlorine was determined in the Cryoem Structure of the Histamine H1 Receptor-Bril/Anti Bril Fab Complex with Desloratadine, PDB code: 8x64:
In total only one binding site of Chlorine was determined in the Cryoem Structure of the Histamine H1 Receptor-Bril/Anti Bril Fab Complex with Desloratadine, PDB code: 8x64:
Chlorine binding site 1 out of 1 in 8x64
Go back to
Chlorine binding site 1 out
of 1 in the Cryoem Structure of the Histamine H1 Receptor-Bril/Anti Bril Fab Complex with Desloratadine
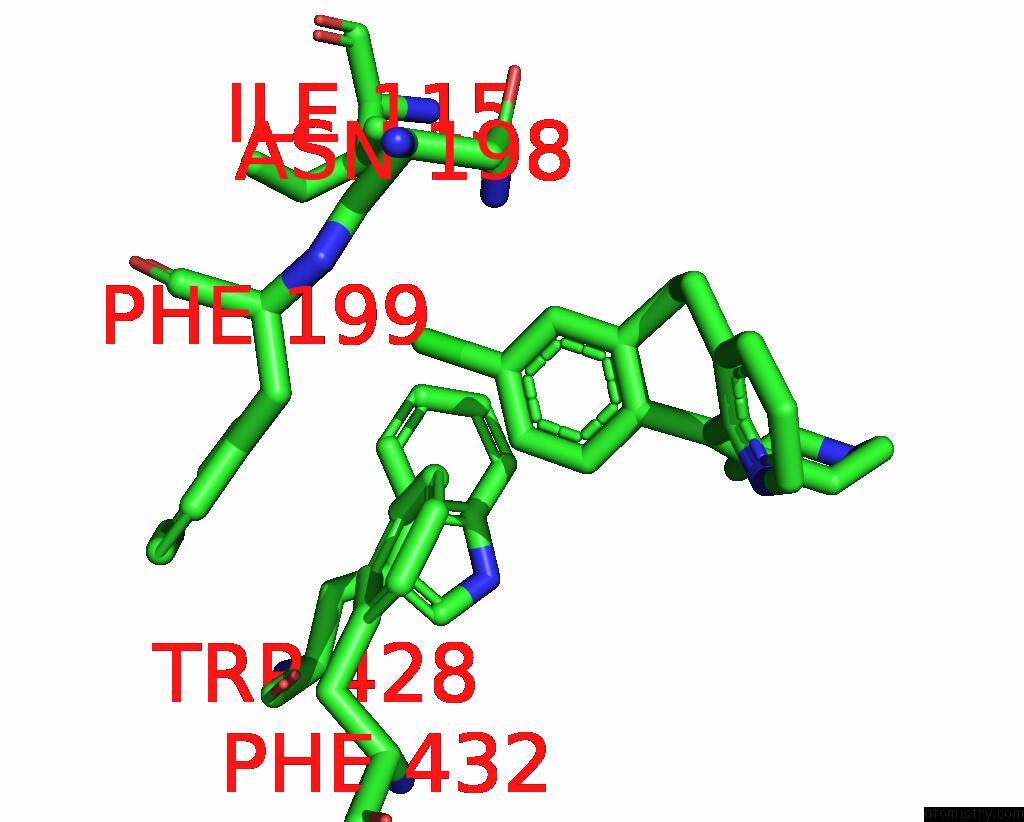
Mono view
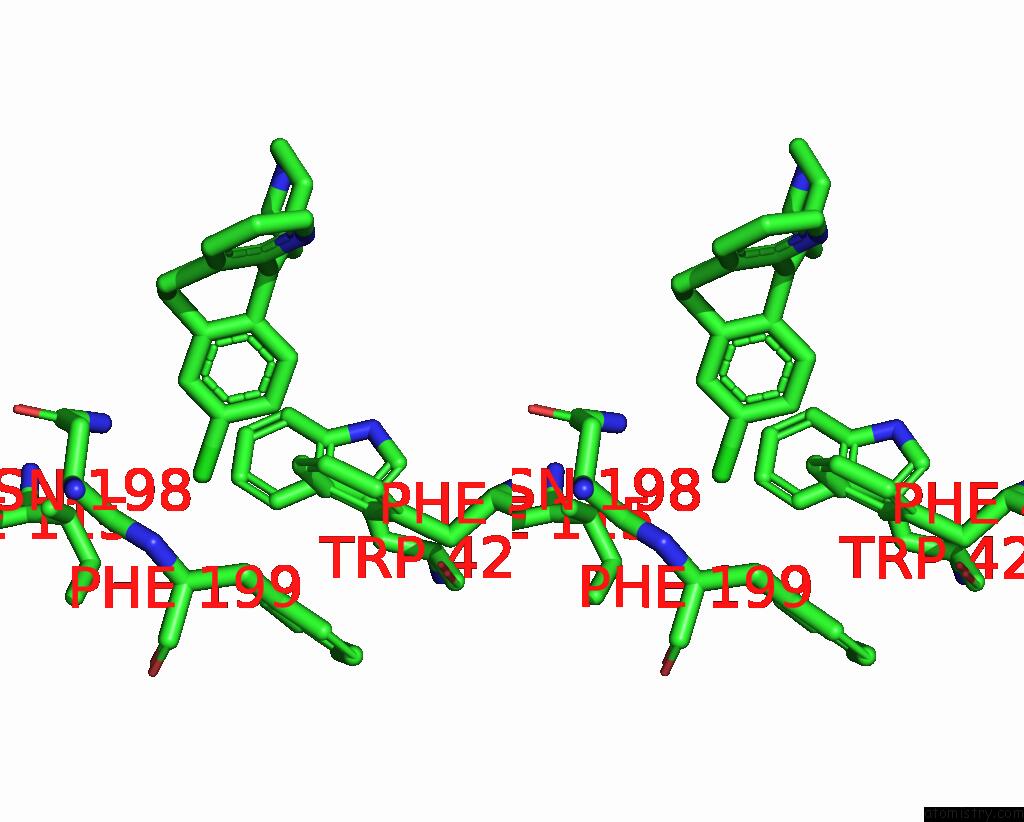
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Cryoem Structure of the Histamine H1 Receptor-Bril/Anti Bril Fab Complex with Desloratadine within 5.0Å range:
|
Reference:
D.Wang,
Q.Guo,
Z.Wu,
M.Li,
B.He,
Y.Du,
K.Zhang,
Y.Tao.
Molecular Mechanism of Antihistamines Recognition and Regulation of the Histamine H 1 Receptor. Nat Commun V. 15 84 2024.
ISSN: ESSN 2041-1723
PubMed: 38167898
DOI: 10.1038/S41467-023-44477-4
Page generated: Sun Jul 13 15:39:49 2025
ISSN: ESSN 2041-1723
PubMed: 38167898
DOI: 10.1038/S41467-023-44477-4
Last articles
F in 7PJCF in 7PJ2
F in 7PHN
F in 7PG6
F in 7PHJ
F in 7PAV
F in 7PH1
F in 7PCD
F in 7P80
F in 7PAW