Chlorine »
PDB 2xev-2xmc »
2xiw »
Chlorine in PDB 2xiw: Crystal Structure of the SAC7D-Derived IGG1-Binder C3-C24S
Protein crystallography data
The structure of Crystal Structure of the SAC7D-Derived IGG1-Binder C3-C24S, PDB code: 2xiw
was solved by
M.Bellinzoni,
S.Colinet,
G.Behar,
P.M.Alzari,
F.Pecorari,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 18.37 / 1.50 |
| Space group | P 31 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 73.175, 73.175, 57.445, 90.00, 90.00, 120.00 |
| R / Rfree (%) | 17.52 / 19.1 |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Crystal Structure of the SAC7D-Derived IGG1-Binder C3-C24S
(pdb code 2xiw). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total only one binding site of Chlorine was determined in the Crystal Structure of the SAC7D-Derived IGG1-Binder C3-C24S, PDB code: 2xiw:
In total only one binding site of Chlorine was determined in the Crystal Structure of the SAC7D-Derived IGG1-Binder C3-C24S, PDB code: 2xiw:
Chlorine binding site 1 out of 1 in 2xiw
Go back to
Chlorine binding site 1 out
of 1 in the Crystal Structure of the SAC7D-Derived IGG1-Binder C3-C24S
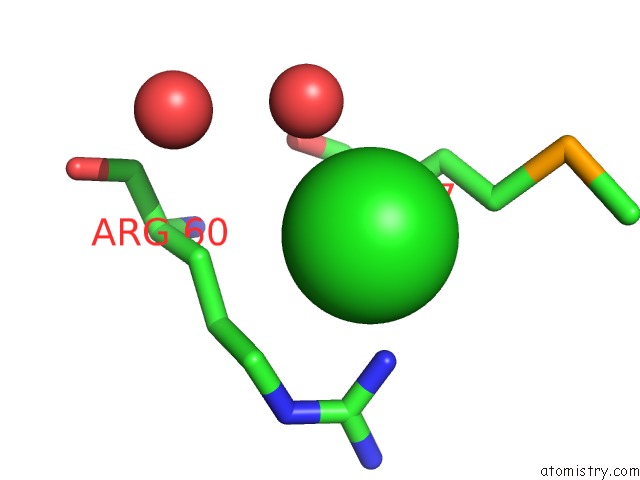
Mono view
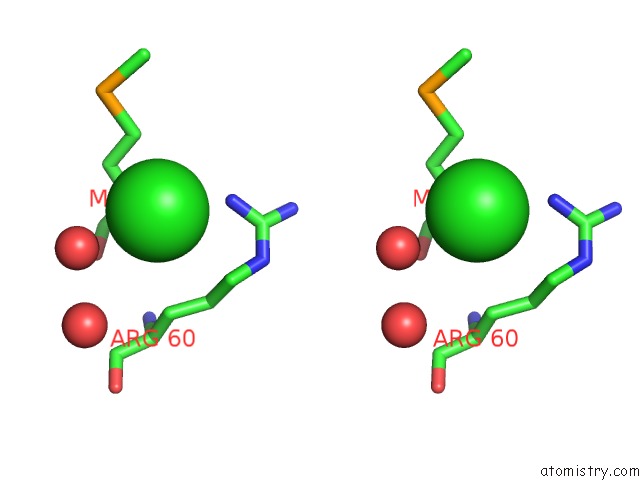
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Crystal Structure of the SAC7D-Derived IGG1-Binder C3-C24S within 5.0Å range:
|
Reference:
G.Behar,
M.Bellinzoni,
M.Maillasson,
L.Paillard-Laurance,
P.M.Alzari,
X.He,
B.Mouratou,
F.Pecorari.
Tolerance of the Archaeal SAC7D Scaffold Protein to Alternative Library Designs: Characterization of Anti- Immunoglobulin G Affitins. Protein Eng.Des.Sel. V. 26 267 2013.
ISSN: ISSN 1741-0126
PubMed: 23315487
DOI: 10.1093/PROTEIN/GZS106
Page generated: Fri Jul 11 01:58:08 2025
ISSN: ISSN 1741-0126
PubMed: 23315487
DOI: 10.1093/PROTEIN/GZS106
Last articles
Na in 2WNTNa in 2WNH
Na in 2WNN
Na in 2WL4
Na in 2WMZ
Na in 2WLU
Na in 2WKV
Na in 2WL5
Na in 2WIS
Na in 2WKT