Chlorine »
PDB 4bbz-4bjy »
4bel »
Chlorine in PDB 4bel: BACE2 Xaperone Complex
Enzymatic activity of BACE2 Xaperone Complex
All present enzymatic activity of BACE2 Xaperone Complex:
3.4.23.45;
3.4.23.45;
Protein crystallography data
The structure of BACE2 Xaperone Complex, PDB code: 4bel
was solved by
D.W.Banner,
A.Kuglstatter,
J.Benz,
M.Stihle,
A.Ruf,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 43.68 / 1.85 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 63.027, 61.757, 115.620, 90.00, 101.38, 90.00 |
| R / Rfree (%) | 18.13 / 22.234 |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the BACE2 Xaperone Complex
(pdb code 4bel). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total only one binding site of Chlorine was determined in the BACE2 Xaperone Complex, PDB code: 4bel:
In total only one binding site of Chlorine was determined in the BACE2 Xaperone Complex, PDB code: 4bel:
Chlorine binding site 1 out of 1 in 4bel
Go back to
Chlorine binding site 1 out
of 1 in the BACE2 Xaperone Complex
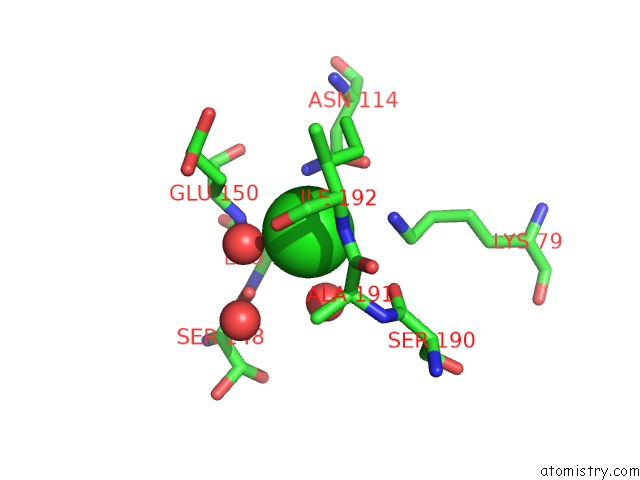
Mono view
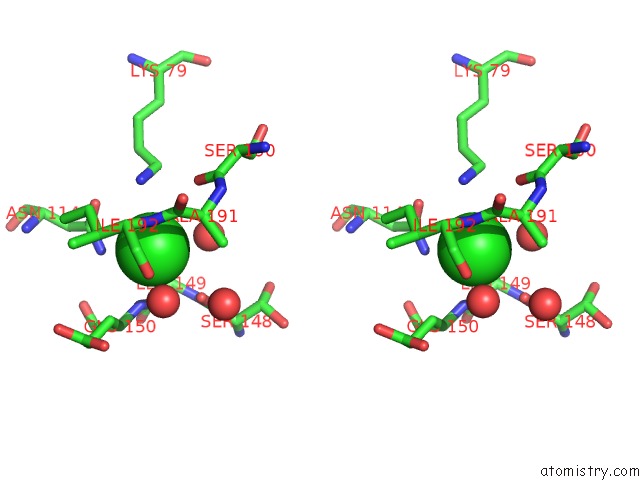
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of BACE2 Xaperone Complex within 5.0Å range:
|
Reference:
D.W.Banner,
B.Gsell,
J.Benz,
J.Bertschinger,
D.Burger,
S.Brack,
S.Cuppuleri,
M.Debulpaep,
A.Gast,
D.Grabulovski,
M.Hennig,
H.Hilpert,
W.Huber,
A.Kuglstatter,
E.Kusznir,
T.Laeremans,
H.Matile,
C.Miscenic,
A.Rufer,
D.Schlatter,
J.Steyeart,
M.Stihle,
R.Thoma,
M.Weber,
A.Ruf.
Mapping the Conformational Space Accessible to BACE2 Using Surface Mutants and Co-Crystals with Fab-Fragments, Fynomers, and Xaperones Acta Crystallogr.,Sect.D V. 69 1124 2013.
ISSN: ISSN 0907-4449
PubMed: 23695257
DOI: 10.1107/S0907444913006574
Page generated: Fri Jul 11 13:19:35 2025
ISSN: ISSN 0907-4449
PubMed: 23695257
DOI: 10.1107/S0907444913006574
Last articles
Mg in 1VQ9Mg in 1VQ7
Mg in 1VQ5
Mg in 1VQ6
Mg in 1VQ8
Mg in 1VQ4
Mg in 1VPA
Mg in 1VPE
Mg in 1VOM
Mg in 1VMA