Chlorine »
PDB 4rsh-4s27 »
4rw9 »
Chlorine in PDB 4rw9: Crystal Structure of Hiv-1 Reverse Transcriptase (Y181C) Variant in Complex with (E)-3-(3-Chloro-5-(2-(2-(2,4-Dioxo-3,4-Dihydropyrimidin- 1(2H)-Yl)Ethoxy)Phenoxy)Phenyl)Acrylonitrile (JLJ532), A Non- Nucleoside Inhibitor
Enzymatic activity of Crystal Structure of Hiv-1 Reverse Transcriptase (Y181C) Variant in Complex with (E)-3-(3-Chloro-5-(2-(2-(2,4-Dioxo-3,4-Dihydropyrimidin- 1(2H)-Yl)Ethoxy)Phenoxy)Phenyl)Acrylonitrile (JLJ532), A Non- Nucleoside Inhibitor
All present enzymatic activity of Crystal Structure of Hiv-1 Reverse Transcriptase (Y181C) Variant in Complex with (E)-3-(3-Chloro-5-(2-(2-(2,4-Dioxo-3,4-Dihydropyrimidin- 1(2H)-Yl)Ethoxy)Phenoxy)Phenyl)Acrylonitrile (JLJ532), A Non- Nucleoside Inhibitor:
2.7.7.49; 2.7.7.7; 3.1.13.2; 3.1.26.13;
2.7.7.49; 2.7.7.7; 3.1.13.2; 3.1.26.13;
Protein crystallography data
The structure of Crystal Structure of Hiv-1 Reverse Transcriptase (Y181C) Variant in Complex with (E)-3-(3-Chloro-5-(2-(2-(2,4-Dioxo-3,4-Dihydropyrimidin- 1(2H)-Yl)Ethoxy)Phenoxy)Phenyl)Acrylonitrile (JLJ532), A Non- Nucleoside Inhibitor, PDB code: 4rw9
was solved by
K.M.Frey,
K.S.Anderson,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 34.81 / 2.99 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 161.416, 74.026, 108.102, 90.00, 99.69, 90.00 |
| R / Rfree (%) | 23.2 / 28 |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Crystal Structure of Hiv-1 Reverse Transcriptase (Y181C) Variant in Complex with (E)-3-(3-Chloro-5-(2-(2-(2,4-Dioxo-3,4-Dihydropyrimidin- 1(2H)-Yl)Ethoxy)Phenoxy)Phenyl)Acrylonitrile (JLJ532), A Non- Nucleoside Inhibitor
(pdb code 4rw9). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total only one binding site of Chlorine was determined in the Crystal Structure of Hiv-1 Reverse Transcriptase (Y181C) Variant in Complex with (E)-3-(3-Chloro-5-(2-(2-(2,4-Dioxo-3,4-Dihydropyrimidin- 1(2H)-Yl)Ethoxy)Phenoxy)Phenyl)Acrylonitrile (JLJ532), A Non- Nucleoside Inhibitor, PDB code: 4rw9:
In total only one binding site of Chlorine was determined in the Crystal Structure of Hiv-1 Reverse Transcriptase (Y181C) Variant in Complex with (E)-3-(3-Chloro-5-(2-(2-(2,4-Dioxo-3,4-Dihydropyrimidin- 1(2H)-Yl)Ethoxy)Phenoxy)Phenyl)Acrylonitrile (JLJ532), A Non- Nucleoside Inhibitor, PDB code: 4rw9:
Chlorine binding site 1 out of 1 in 4rw9
Go back to
Chlorine binding site 1 out
of 1 in the Crystal Structure of Hiv-1 Reverse Transcriptase (Y181C) Variant in Complex with (E)-3-(3-Chloro-5-(2-(2-(2,4-Dioxo-3,4-Dihydropyrimidin- 1(2H)-Yl)Ethoxy)Phenoxy)Phenyl)Acrylonitrile (JLJ532), A Non- Nucleoside Inhibitor
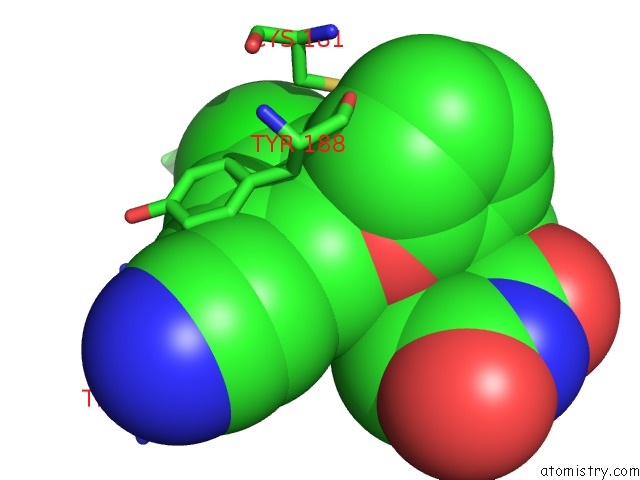
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Crystal Structure of Hiv-1 Reverse Transcriptase (Y181C) Variant in Complex with (E)-3-(3-Chloro-5-(2-(2-(2,4-Dioxo-3,4-Dihydropyrimidin- 1(2H)-Yl)Ethoxy)Phenoxy)Phenyl)Acrylonitrile (JLJ532), A Non- Nucleoside Inhibitor within 5.0Å range:
|
Reference:
K.M.Frey,
D.E.Puleo,
K.A.Spasov,
M.Bollini,
W.L.Jorgensen,
K.S.Anderson.
Structure-Based Evaluation of Non-Nucleoside Inhibitors with Improved Potency and Solubility That Target Hiv Reverse Transcriptase Variants. J.Med.Chem. V. 58 2737 2015.
ISSN: ISSN 0022-2623
PubMed: 25700160
DOI: 10.1021/JM501908A
Page generated: Fri Jul 11 21:37:08 2025
ISSN: ISSN 0022-2623
PubMed: 25700160
DOI: 10.1021/JM501908A
Last articles
Na in 2JBWNa in 2JC4
Na in 2JBY
Na in 2JBA
Na in 2J9Y
Na in 2J9Z
Na in 2J9N
Na in 2J80
Na in 2J9E
Na in 2J5W