Chlorine »
PDB 5ivs-5j7f »
5iwi »
Chlorine in PDB 5iwi: 1.98A Structure of GSK945237 with S.Aureus Dna Gyrase and Singly Nicked Dna
Enzymatic activity of 1.98A Structure of GSK945237 with S.Aureus Dna Gyrase and Singly Nicked Dna
All present enzymatic activity of 1.98A Structure of GSK945237 with S.Aureus Dna Gyrase and Singly Nicked Dna:
5.99.1.3;
5.99.1.3;
Protein crystallography data
The structure of 1.98A Structure of GSK945237 with S.Aureus Dna Gyrase and Singly Nicked Dna, PDB code: 5iwi
was solved by
B.D.Bax,
T.J.Miles,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 18.00 / 1.98 |
| Space group | P 61 |
| Cell size a, b, c (Å), α, β, γ (°) | 92.897, 92.897, 410.784, 90.00, 90.00, 120.00 |
| R / Rfree (%) | 16.6 / 20.2 |
Other elements in 5iwi:
The structure of 1.98A Structure of GSK945237 with S.Aureus Dna Gyrase and Singly Nicked Dna also contains other interesting chemical elements:
| Fluorine | (F) | 2 atoms |
| Manganese | (Mn) | 5 atoms |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the 1.98A Structure of GSK945237 with S.Aureus Dna Gyrase and Singly Nicked Dna
(pdb code 5iwi). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total only one binding site of Chlorine was determined in the 1.98A Structure of GSK945237 with S.Aureus Dna Gyrase and Singly Nicked Dna, PDB code: 5iwi:
In total only one binding site of Chlorine was determined in the 1.98A Structure of GSK945237 with S.Aureus Dna Gyrase and Singly Nicked Dna, PDB code: 5iwi:
Chlorine binding site 1 out of 1 in 5iwi
Go back to
Chlorine binding site 1 out
of 1 in the 1.98A Structure of GSK945237 with S.Aureus Dna Gyrase and Singly Nicked Dna
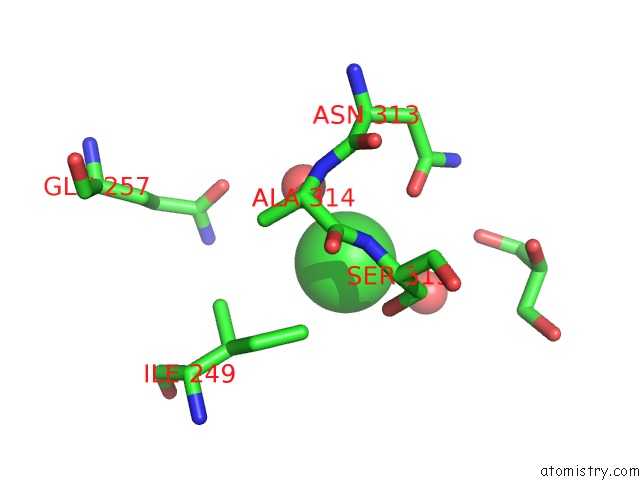
Mono view
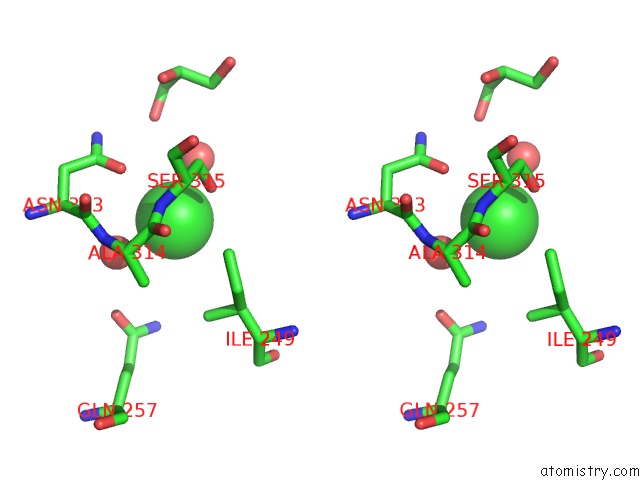
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of 1.98A Structure of GSK945237 with S.Aureus Dna Gyrase and Singly Nicked Dna within 5.0Å range:
|
Reference:
T.J.Miles,
A.J.Hennessy,
B.Bax,
G.Brooks,
B.S.Brown,
P.Brown,
N.Cailleau,
D.Chen,
S.Dabbs,
D.T.Davies,
J.M.Esken,
I.Giordano,
J.L.Hoover,
G.E.Jones,
S.K.Kusalakumari Sukmar,
R.E.Markwell,
E.A.Minthorn,
S.Rittenhouse,
M.N.Gwynn,
N.D.Pearson.
Novel Tricyclics (E.G., GSK945237) As Potent Inhibitors of Bacterial Type Iia Topoisomerases. Bioorg.Med.Chem.Lett. V. 26 2464 2016.
ISSN: ESSN 1464-3405
PubMed: 27055939
DOI: 10.1016/J.BMCL.2016.03.106
Page generated: Sat Jul 12 03:19:36 2025
ISSN: ESSN 1464-3405
PubMed: 27055939
DOI: 10.1016/J.BMCL.2016.03.106
Last articles
Mg in 9MF6Mg in 9MED
Mg in 9MEC
Mg in 9MDY
Mg in 9MDW
Mg in 9M84
Mg in 9M7M
Mg in 9M7L
Mg in 9M5O
Mg in 9M56