Chlorine »
PDB 5t79-5ti4 »
5t9p »
Chlorine in PDB 5t9p: Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate
Protein crystallography data
The structure of Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate, PDB code: 5t9p
was solved by
J.Zhang,
E.L.Gonzalez,
M.T.T.Hall,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 45.00 / 2.00 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 37.242, 157.399, 47.130, 90.00, 107.06, 90.00 |
| R / Rfree (%) | 19.2 / 23.6 |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate
(pdb code 5t9p). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total 7 binding sites of Chlorine where determined in the Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate, PDB code: 5t9p:
Jump to Chlorine binding site number: 1; 2; 3; 4; 5; 6; 7;
In total 7 binding sites of Chlorine where determined in the Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate, PDB code: 5t9p:
Jump to Chlorine binding site number: 1; 2; 3; 4; 5; 6; 7;
Chlorine binding site 1 out of 7 in 5t9p
Go back to
Chlorine binding site 1 out
of 7 in the Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate
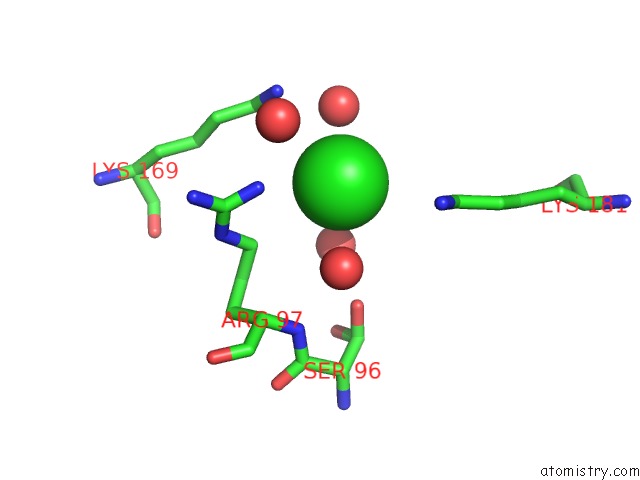
Mono view
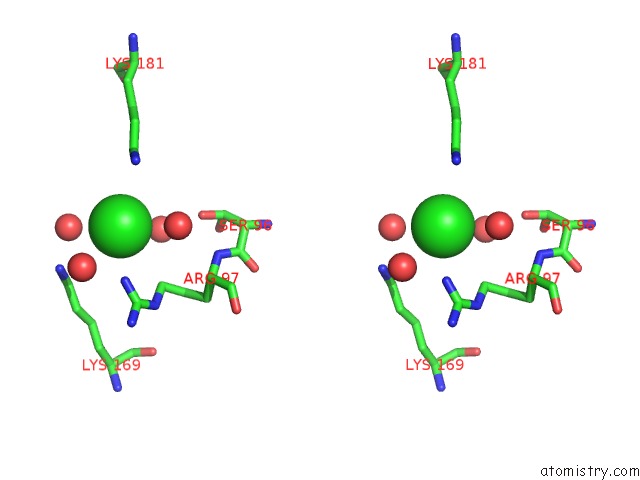
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate within 5.0Å range:
|
Chlorine binding site 2 out of 7 in 5t9p
Go back to
Chlorine binding site 2 out
of 7 in the Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate
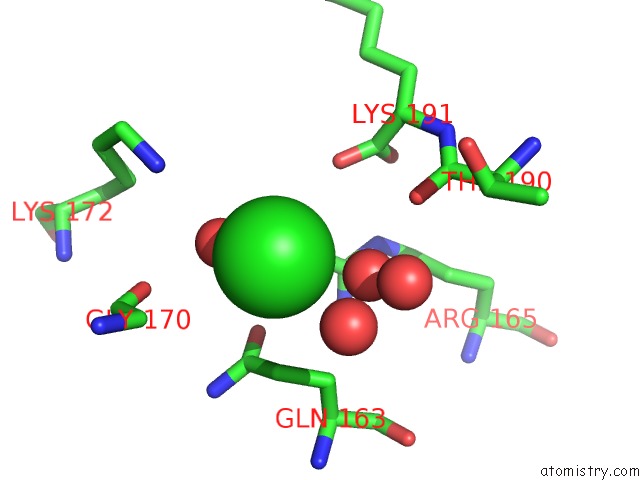
Mono view
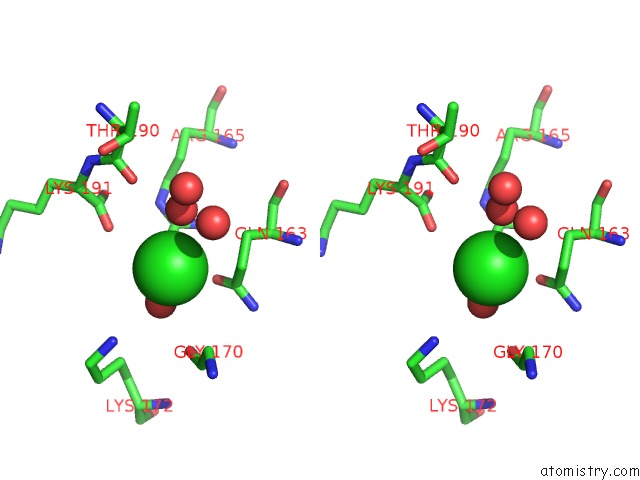
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 2 of Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate within 5.0Å range:
|
Chlorine binding site 3 out of 7 in 5t9p
Go back to
Chlorine binding site 3 out
of 7 in the Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate
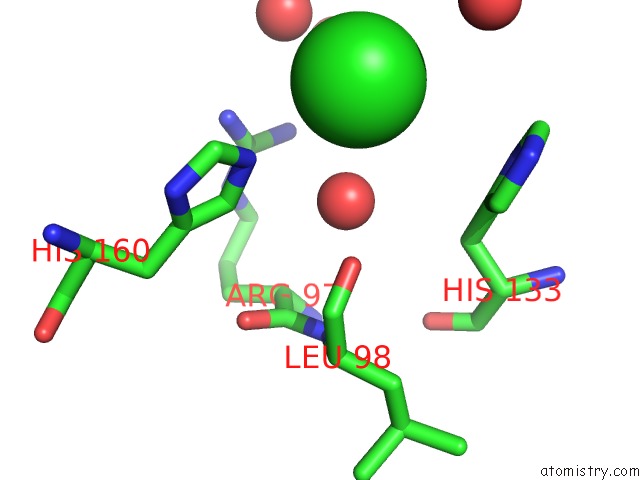
Mono view
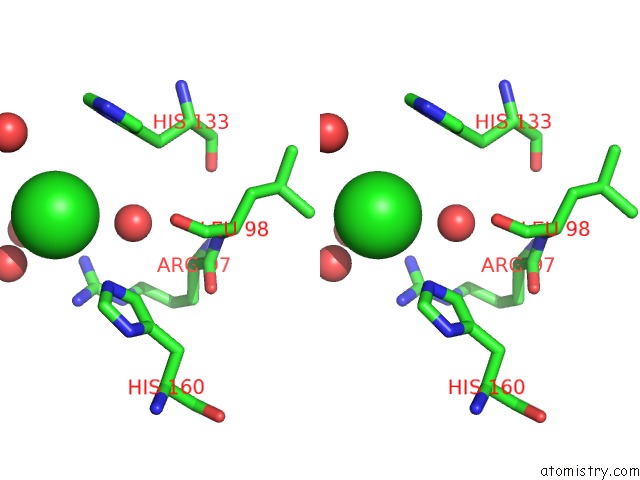
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 3 of Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate within 5.0Å range:
|
Chlorine binding site 4 out of 7 in 5t9p
Go back to
Chlorine binding site 4 out
of 7 in the Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate
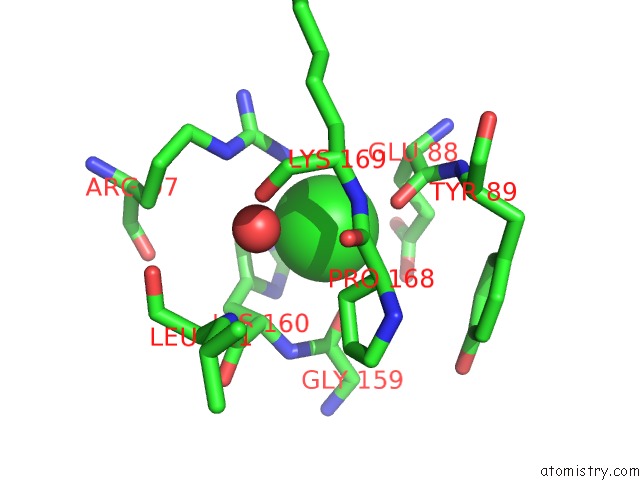
Mono view
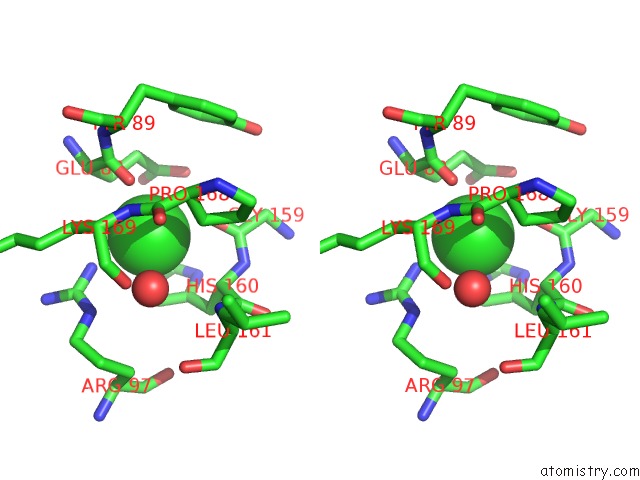
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 4 of Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate within 5.0Å range:
|
Chlorine binding site 5 out of 7 in 5t9p
Go back to
Chlorine binding site 5 out
of 7 in the Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate
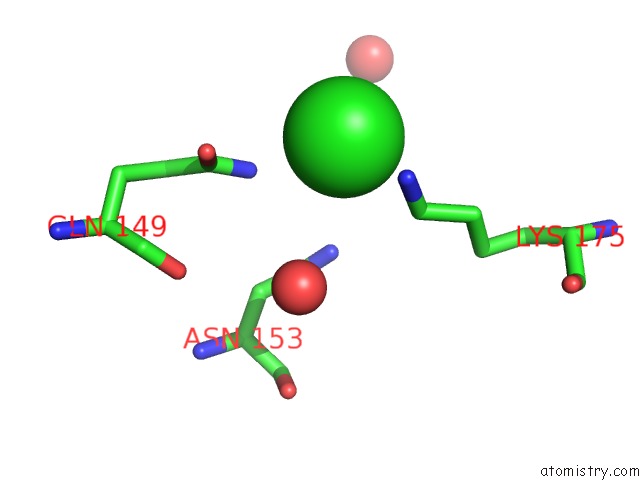
Mono view
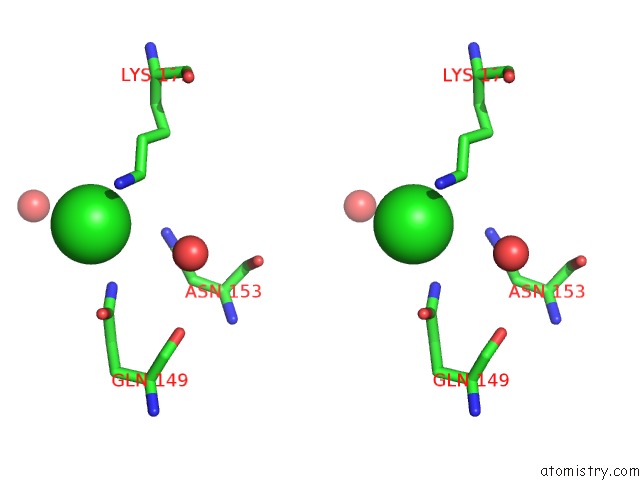
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 5 of Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate within 5.0Å range:
|
Chlorine binding site 6 out of 7 in 5t9p
Go back to
Chlorine binding site 6 out
of 7 in the Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate
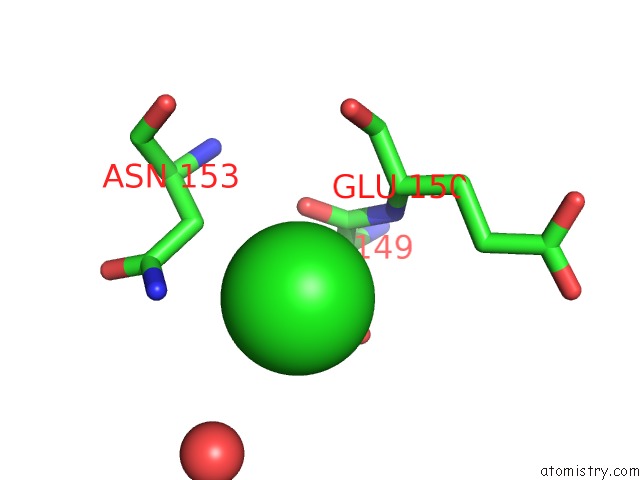
Mono view
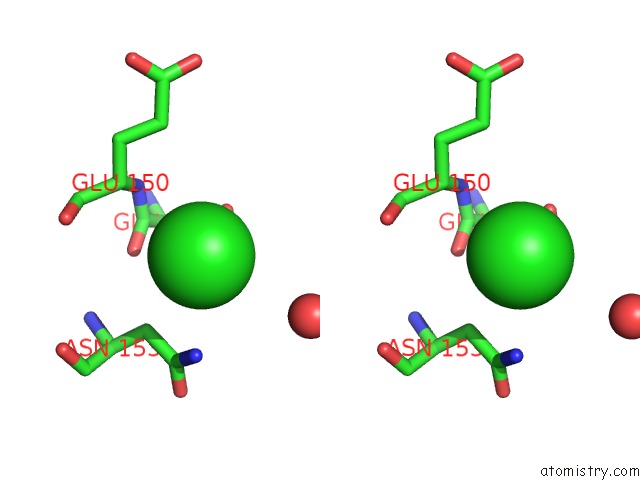
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 6 of Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate within 5.0Å range:
|
Chlorine binding site 7 out of 7 in 5t9p
Go back to
Chlorine binding site 7 out
of 7 in the Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate
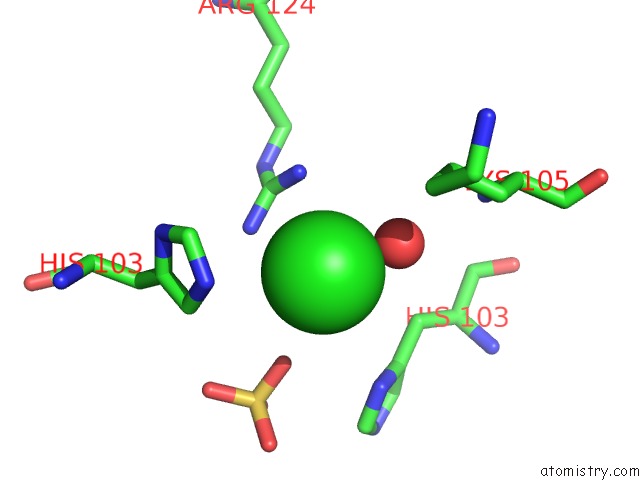
Mono view
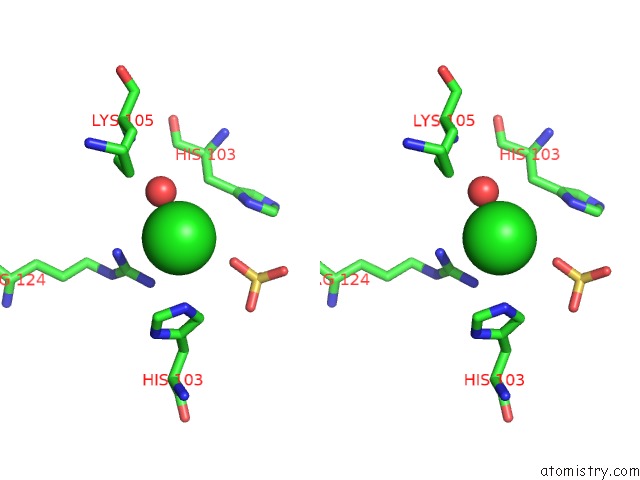
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 7 of Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate within 5.0Å range:
|
Reference:
J.Zhang,
L.E.Gonzalez,
T.M.T.Hall.
Structural Analysis Reveals the Flexible C-Terminus of NOP15 Undergoes Rearrangement to Recognize A Pre-Ribosomal Rna Folding Intermediate. Nucleic Acids Res. V. 45 2829 2017.
ISSN: ESSN 1362-4962
PubMed: 27789691
DOI: 10.1093/NAR/GKW961
Page generated: Sat Jul 12 08:51:18 2025
ISSN: ESSN 1362-4962
PubMed: 27789691
DOI: 10.1093/NAR/GKW961
Last articles
Zn in 1XF7Zn in 1XER
Zn in 1XEM
Zn in 1XEG
Zn in 1XDJ
Zn in 1XDA
Zn in 1XCR
Zn in 1XBT
Zn in 1XC8
Zn in 1XC3