Chlorine »
PDB 4ibf-4iij »
4idg »
Chlorine in PDB 4idg: Crystal Structure of A Short-Chain Dehydrogenase/Reductase Superfamily Protein From Agrobacterium Tumefaciens (Target Efi-506441) with Bound Nad, Monoclinic Form 2
Protein crystallography data
The structure of Crystal Structure of A Short-Chain Dehydrogenase/Reductase Superfamily Protein From Agrobacterium Tumefaciens (Target Efi-506441) with Bound Nad, Monoclinic Form 2, PDB code: 4idg
was solved by
M.W.Vetting,
F.Groninger-Poe,
L.L.Morisco,
S.R.Wasserman,
S.Sojitra,
H.J.Imker,
J.A.Gerlt,
S.C.Almo,
Enzyme Function Initiative (Efi),
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 35.75 / 1.90 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 77.388, 57.703, 95.418, 90.00, 108.68, 90.00 |
| R / Rfree (%) | 17.7 / 21.6 |
Chlorine Binding Sites:
The binding sites of Chlorine atom in the Crystal Structure of A Short-Chain Dehydrogenase/Reductase Superfamily Protein From Agrobacterium Tumefaciens (Target Efi-506441) with Bound Nad, Monoclinic Form 2
(pdb code 4idg). This binding sites where shown within
5.0 Angstroms radius around Chlorine atom.
In total 3 binding sites of Chlorine where determined in the Crystal Structure of A Short-Chain Dehydrogenase/Reductase Superfamily Protein From Agrobacterium Tumefaciens (Target Efi-506441) with Bound Nad, Monoclinic Form 2, PDB code: 4idg:
Jump to Chlorine binding site number: 1; 2; 3;
In total 3 binding sites of Chlorine where determined in the Crystal Structure of A Short-Chain Dehydrogenase/Reductase Superfamily Protein From Agrobacterium Tumefaciens (Target Efi-506441) with Bound Nad, Monoclinic Form 2, PDB code: 4idg:
Jump to Chlorine binding site number: 1; 2; 3;
Chlorine binding site 1 out of 3 in 4idg
Go back to
Chlorine binding site 1 out
of 3 in the Crystal Structure of A Short-Chain Dehydrogenase/Reductase Superfamily Protein From Agrobacterium Tumefaciens (Target Efi-506441) with Bound Nad, Monoclinic Form 2
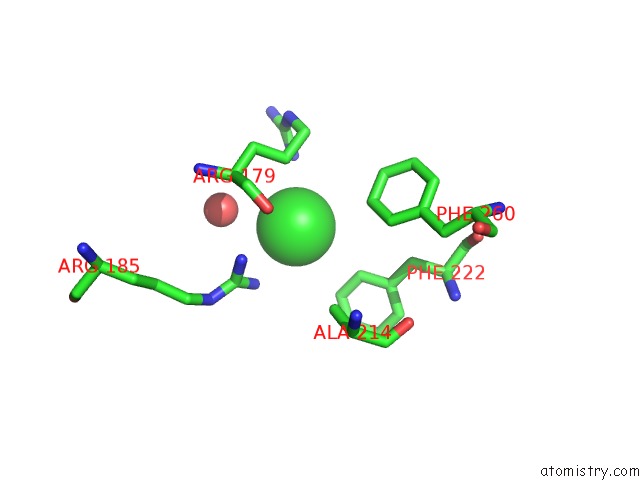
Mono view
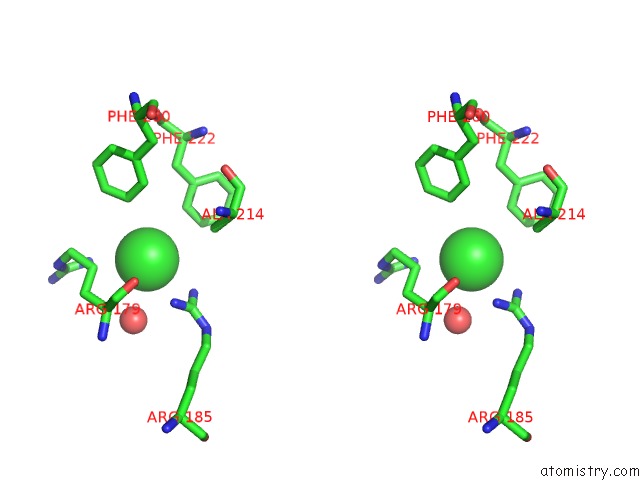
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of Crystal Structure of A Short-Chain Dehydrogenase/Reductase Superfamily Protein From Agrobacterium Tumefaciens (Target Efi-506441) with Bound Nad, Monoclinic Form 2 within 5.0Å range:
|
Chlorine binding site 2 out of 3 in 4idg
Go back to
Chlorine binding site 2 out
of 3 in the Crystal Structure of A Short-Chain Dehydrogenase/Reductase Superfamily Protein From Agrobacterium Tumefaciens (Target Efi-506441) with Bound Nad, Monoclinic Form 2
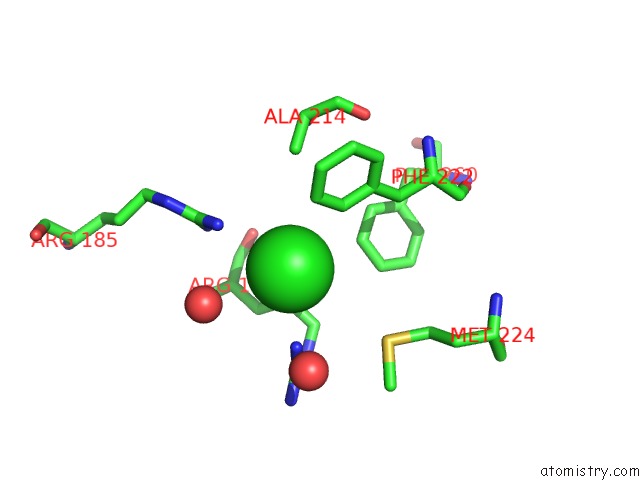
Mono view
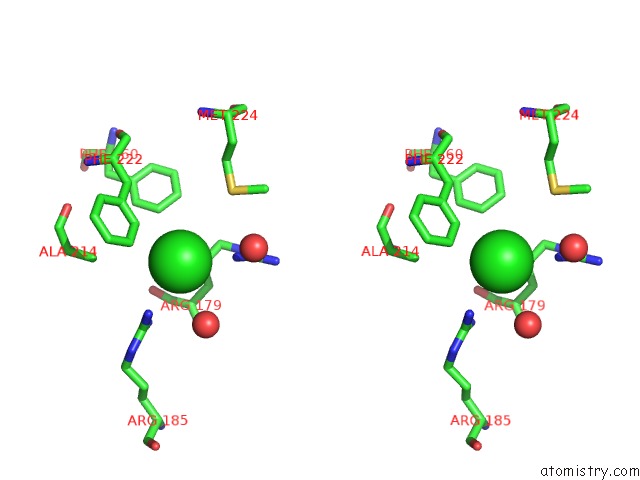
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 2 of Crystal Structure of A Short-Chain Dehydrogenase/Reductase Superfamily Protein From Agrobacterium Tumefaciens (Target Efi-506441) with Bound Nad, Monoclinic Form 2 within 5.0Å range:
|
Chlorine binding site 3 out of 3 in 4idg
Go back to
Chlorine binding site 3 out
of 3 in the Crystal Structure of A Short-Chain Dehydrogenase/Reductase Superfamily Protein From Agrobacterium Tumefaciens (Target Efi-506441) with Bound Nad, Monoclinic Form 2
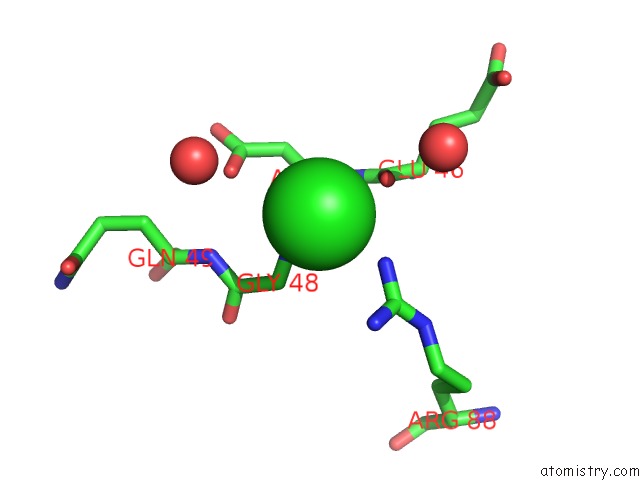
Mono view
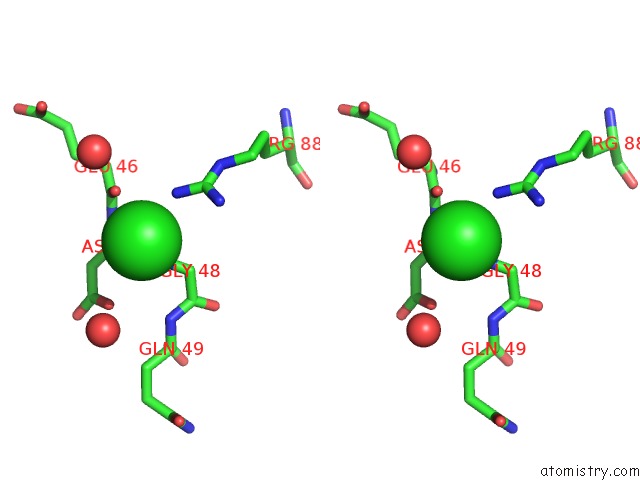
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 3 of Crystal Structure of A Short-Chain Dehydrogenase/Reductase Superfamily Protein From Agrobacterium Tumefaciens (Target Efi-506441) with Bound Nad, Monoclinic Form 2 within 5.0Å range:
|
Reference:
M.W.Vetting,
F.Groninger-Poe,
L.L.Morisco,
S.R.Wasserman,
S.Sojitra,
H.J.Imker,
J.A.Gerlt,
S.C.Almo,
Enzyme Function Initiative (Efi).
Crystal Structure of A Short-Chain Dehydrogenase/Reductase Superfamily Protein From Agrobacterium Tumefaciens (Target Efi-506441) with Bound Nad, Monoclinic Form 2 To Be Published.
Page generated: Fri Jul 11 16:48:14 2025
Last articles
Cl in 4WPICl in 4WPH
Cl in 4WQK
Cl in 4WPL
Cl in 4WP8
Cl in 4WO5
Cl in 4WP3
Cl in 4WP7
Cl in 4WMV
Cl in 4WMU