Chlorine »
PDB 5l5b-5l7f »
5l6i »
Chlorine in PDB 5l6i: UBA1 in Complex with Ub-MLN4924 Covalent Adduct
Enzymatic activity of UBA1 in Complex with Ub-MLN4924 Covalent Adduct
All present enzymatic activity of UBA1 in Complex with Ub-MLN4924 Covalent Adduct:
6.2.1.45;
6.2.1.45;
Protein crystallography data
The structure of UBA1 in Complex with Ub-MLN4924 Covalent Adduct, PDB code: 5l6i
was solved by
M.Misra,
H.Schindelin,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 20.00 / 2.76 |
| Space group | P 2 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 72.500, 191.890, 230.197, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 17.4 / 22 |
Chlorine Binding Sites:
Pages:
>>> Page 1 <<< Page 2, Binding sites: 11 - 16;Binding sites:
The binding sites of Chlorine atom in the UBA1 in Complex with Ub-MLN4924 Covalent Adduct (pdb code 5l6i). This binding sites where shown within 5.0 Angstroms radius around Chlorine atom.In total 16 binding sites of Chlorine where determined in the UBA1 in Complex with Ub-MLN4924 Covalent Adduct, PDB code: 5l6i:
Jump to Chlorine binding site number: 1; 2; 3; 4; 5; 6; 7; 8; 9; 10;
Chlorine binding site 1 out of 16 in 5l6i
Go back to
Chlorine binding site 1 out
of 16 in the UBA1 in Complex with Ub-MLN4924 Covalent Adduct
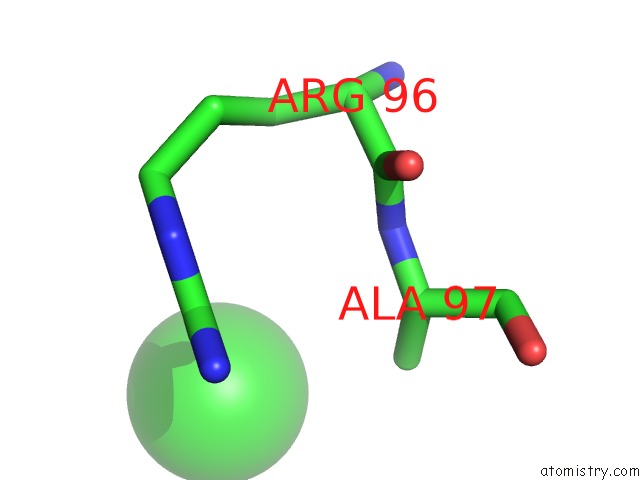
Mono view
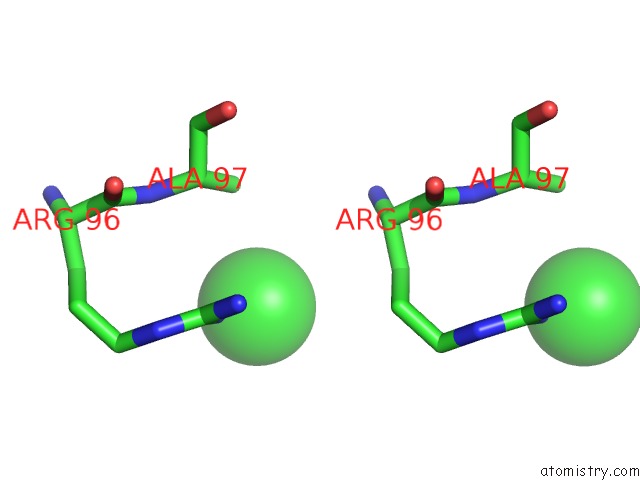
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 1 of UBA1 in Complex with Ub-MLN4924 Covalent Adduct within 5.0Å range:
|
Chlorine binding site 2 out of 16 in 5l6i
Go back to
Chlorine binding site 2 out
of 16 in the UBA1 in Complex with Ub-MLN4924 Covalent Adduct
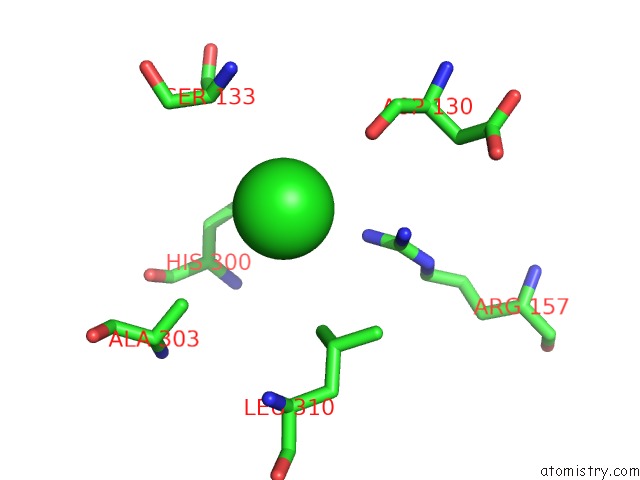
Mono view
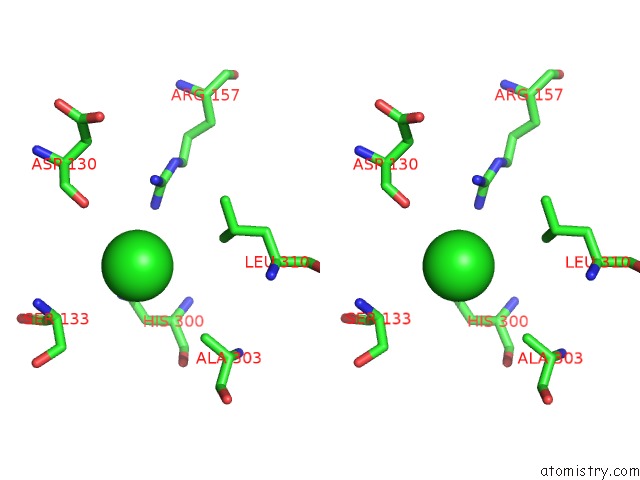
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 2 of UBA1 in Complex with Ub-MLN4924 Covalent Adduct within 5.0Å range:
|
Chlorine binding site 3 out of 16 in 5l6i
Go back to
Chlorine binding site 3 out
of 16 in the UBA1 in Complex with Ub-MLN4924 Covalent Adduct
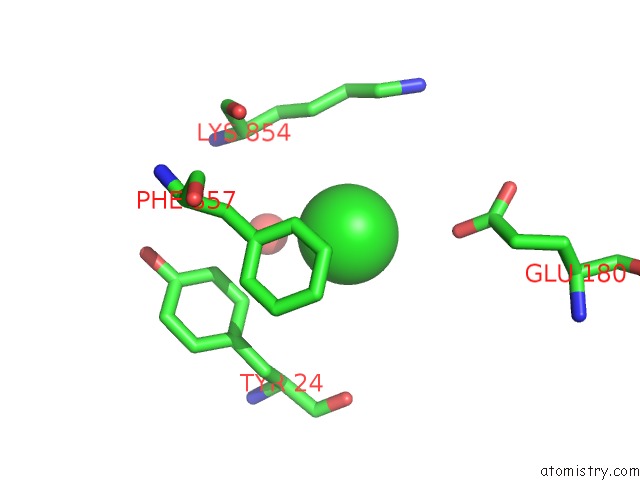
Mono view
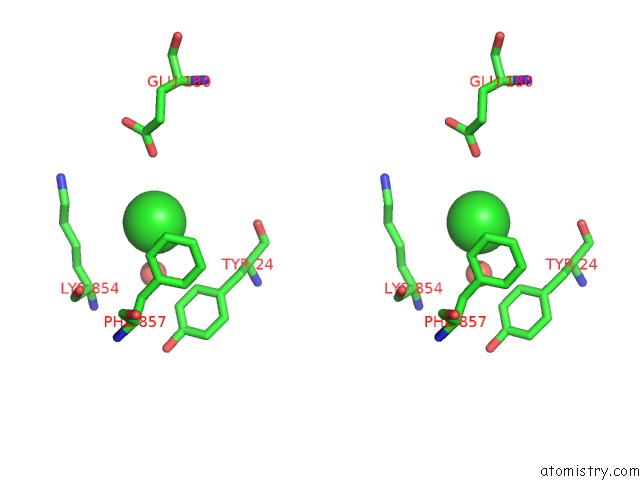
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 3 of UBA1 in Complex with Ub-MLN4924 Covalent Adduct within 5.0Å range:
|
Chlorine binding site 4 out of 16 in 5l6i
Go back to
Chlorine binding site 4 out
of 16 in the UBA1 in Complex with Ub-MLN4924 Covalent Adduct
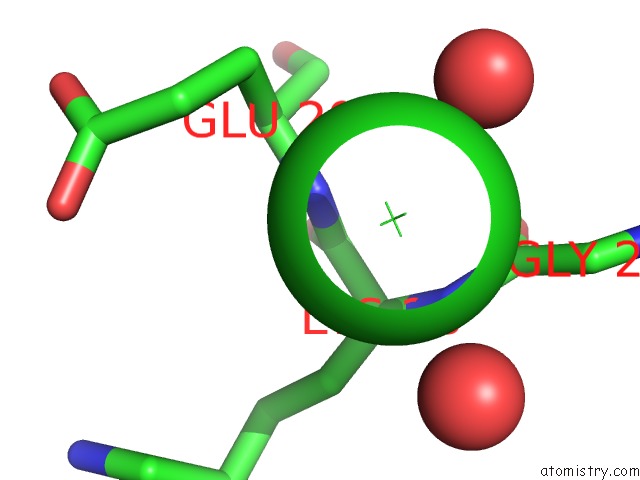
Mono view
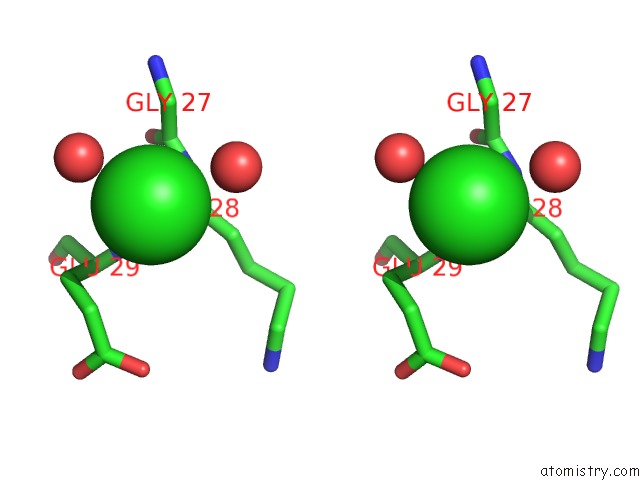
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 4 of UBA1 in Complex with Ub-MLN4924 Covalent Adduct within 5.0Å range:
|
Chlorine binding site 5 out of 16 in 5l6i
Go back to
Chlorine binding site 5 out
of 16 in the UBA1 in Complex with Ub-MLN4924 Covalent Adduct
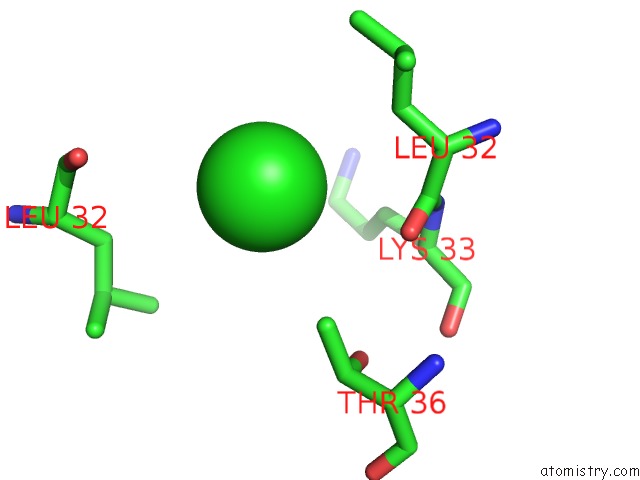
Mono view
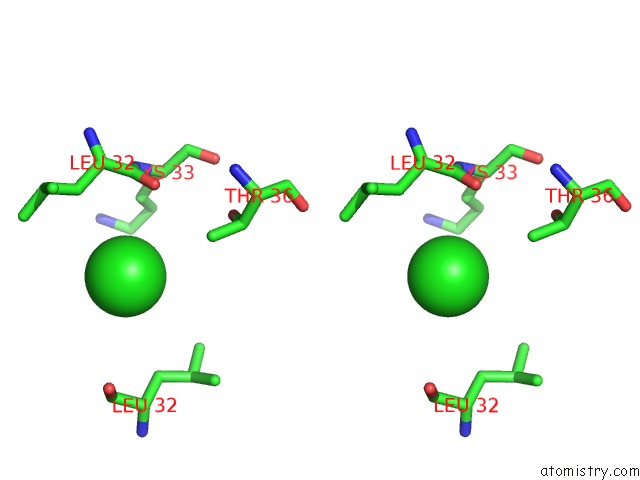
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 5 of UBA1 in Complex with Ub-MLN4924 Covalent Adduct within 5.0Å range:
|
Chlorine binding site 6 out of 16 in 5l6i
Go back to
Chlorine binding site 6 out
of 16 in the UBA1 in Complex with Ub-MLN4924 Covalent Adduct
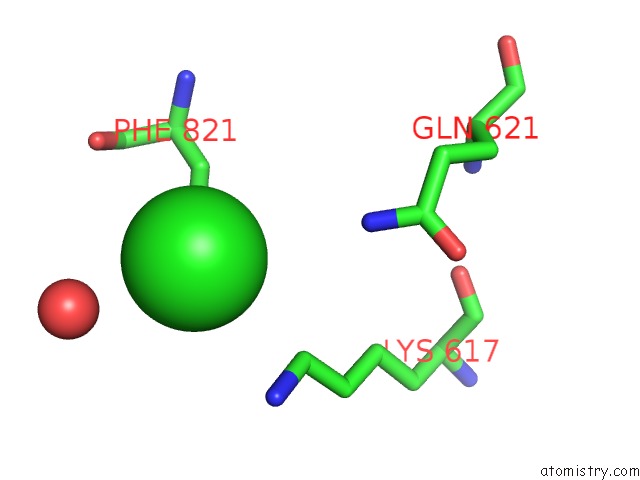
Mono view
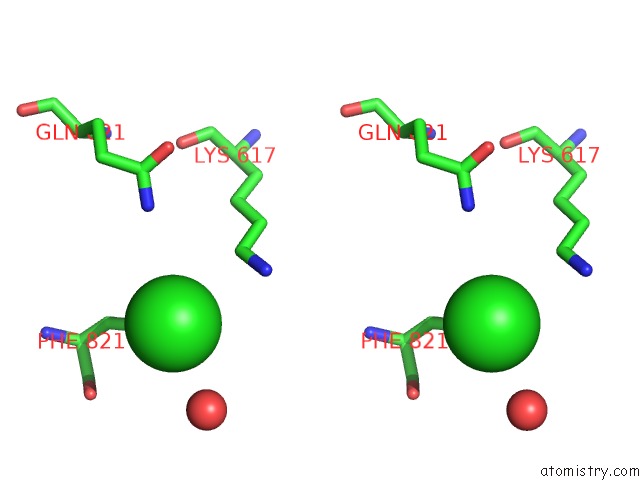
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 6 of UBA1 in Complex with Ub-MLN4924 Covalent Adduct within 5.0Å range:
|
Chlorine binding site 7 out of 16 in 5l6i
Go back to
Chlorine binding site 7 out
of 16 in the UBA1 in Complex with Ub-MLN4924 Covalent Adduct
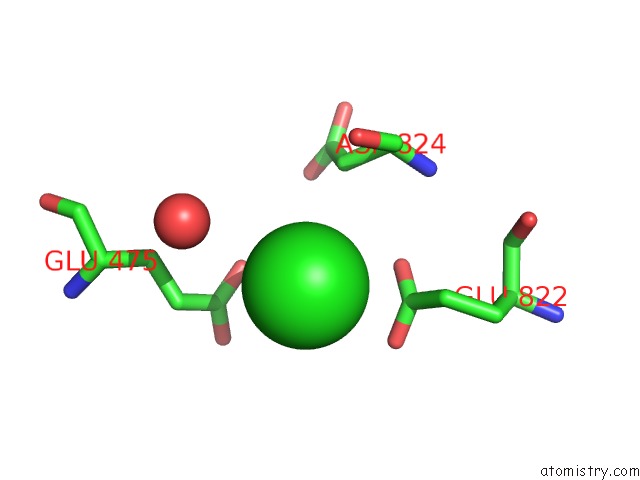
Mono view
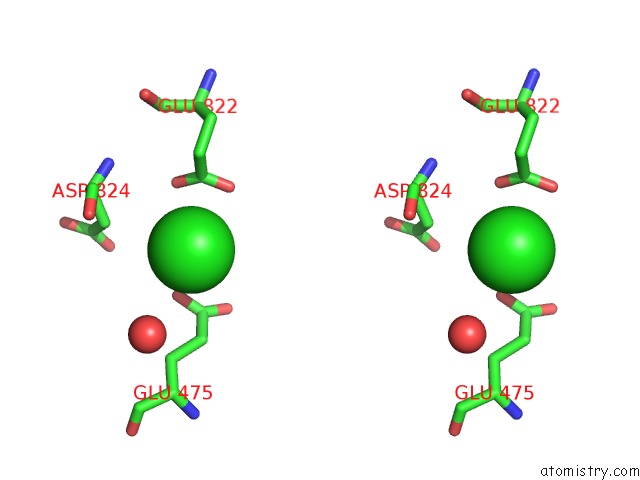
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 7 of UBA1 in Complex with Ub-MLN4924 Covalent Adduct within 5.0Å range:
|
Chlorine binding site 8 out of 16 in 5l6i
Go back to
Chlorine binding site 8 out
of 16 in the UBA1 in Complex with Ub-MLN4924 Covalent Adduct
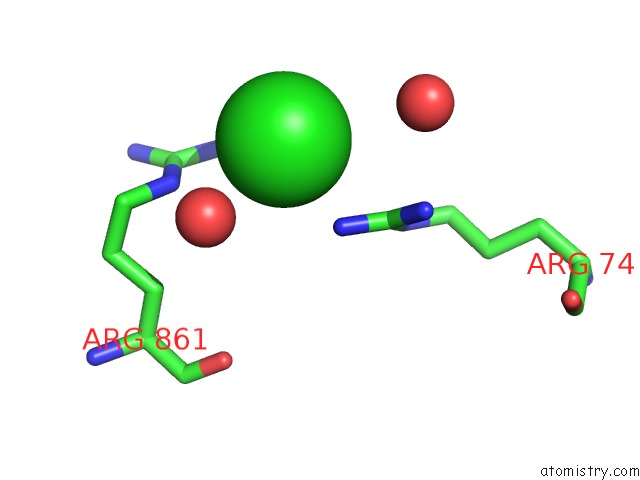
Mono view
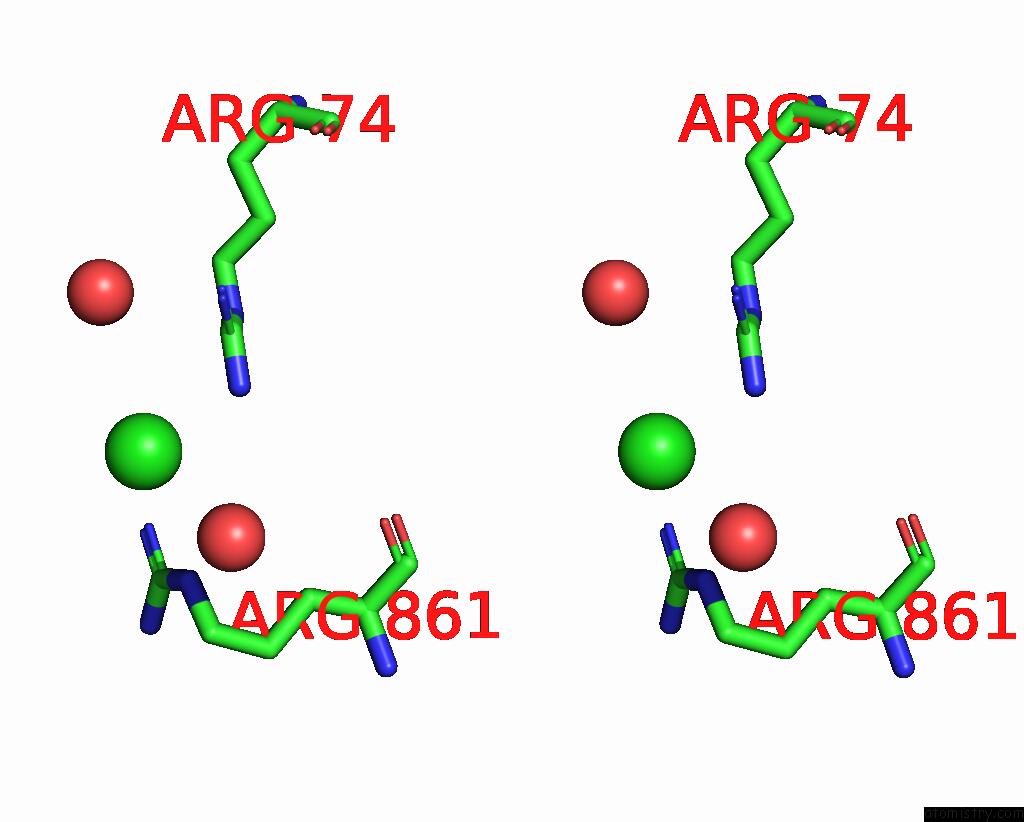
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 8 of UBA1 in Complex with Ub-MLN4924 Covalent Adduct within 5.0Å range:
|
Chlorine binding site 9 out of 16 in 5l6i
Go back to
Chlorine binding site 9 out
of 16 in the UBA1 in Complex with Ub-MLN4924 Covalent Adduct
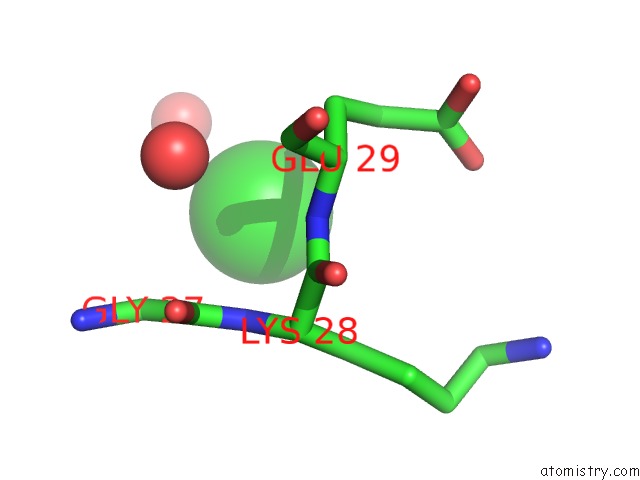
Mono view
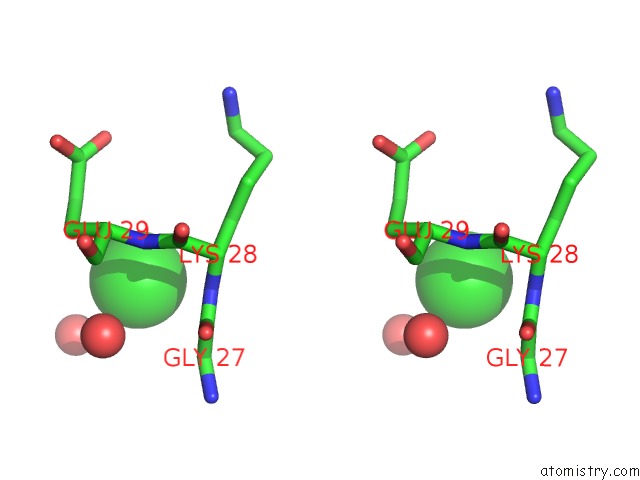
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 9 of UBA1 in Complex with Ub-MLN4924 Covalent Adduct within 5.0Å range:
|
Chlorine binding site 10 out of 16 in 5l6i
Go back to
Chlorine binding site 10 out
of 16 in the UBA1 in Complex with Ub-MLN4924 Covalent Adduct
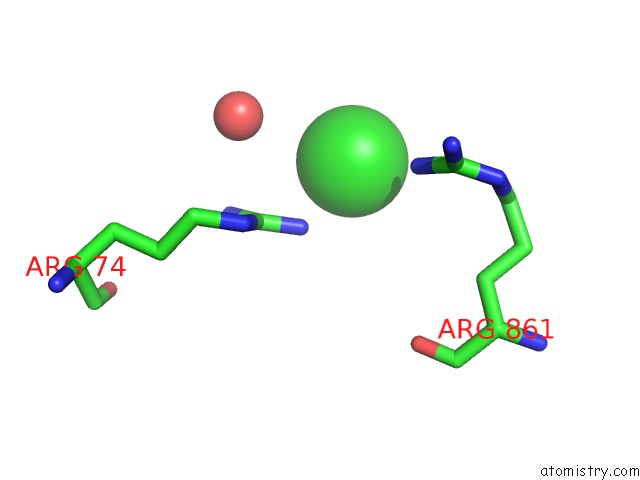
Mono view
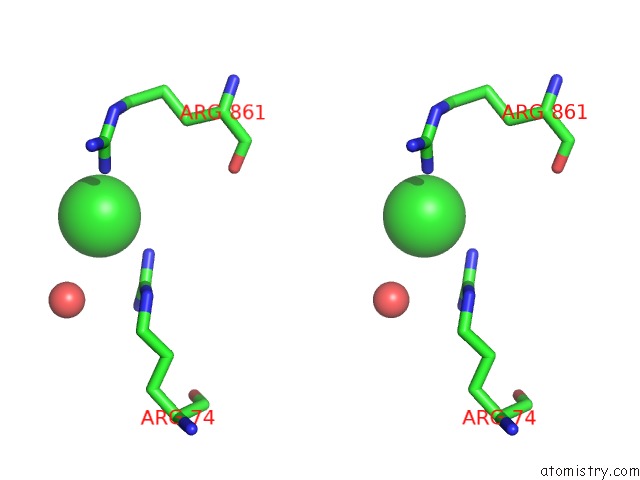
Stereo pair view

Mono view

Stereo pair view
A full contact list of Chlorine with other atoms in the Cl binding
site number 10 of UBA1 in Complex with Ub-MLN4924 Covalent Adduct within 5.0Å range:
|
Reference:
M.Misra,
M.Kuhn,
M.Lobel,
H.An,
A.V.Statsyuk,
C.Sotriffer,
H.Schindelin.
Dissecting the Specificity of Adenosyl Sulfamate Inhibitors Targeting the Ubiquitin-Activating Enzyme. Structure V. 25 1120 2017.
ISSN: ISSN 1878-4186
PubMed: 28578874
DOI: 10.1016/J.STR.2017.05.001
Page generated: Sat Jul 12 04:26:13 2025
ISSN: ISSN 1878-4186
PubMed: 28578874
DOI: 10.1016/J.STR.2017.05.001
Last articles
Cl in 7ZOUCl in 7ZNT
Cl in 7ZO2
Cl in 7ZNP
Cl in 7ZJV
Cl in 7ZLT
Cl in 7ZKS
Cl in 7ZKX
Cl in 7ZKR
Cl in 7ZJU